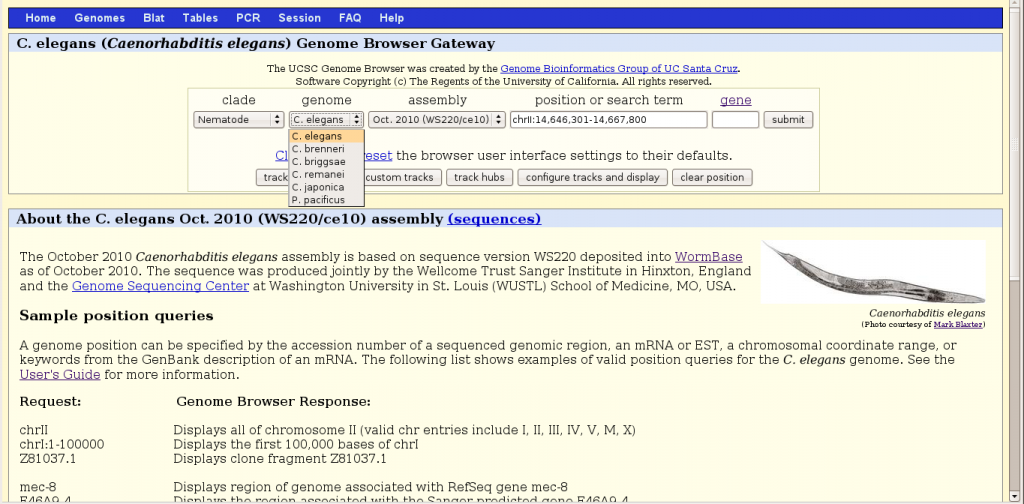
To begin, one should chose the nematode clade then the organism C. elegans, a version of genome sequence assembly (e.g. WS220), and a region of the genome. Browser behavior is context sensitive, dependent on the choices made in the order from left to right. There are three basic ways to specify a region to browse, by a range of chromosomal numerical positions, by a gene name or by an accession number. More flexibly, instead of exact positions, one can also search for a descriptive term (such as “kinase inhibitor”) that is present in gene records. Some restrictions are worth noting. Chromosome positions must start with “chr“. Gene names recognized by the browser appear to be limited to those in RefSeq and Ensembl entries. Remarkably, most if not all named miRNA genes, like lin-4, let-7, lsy-6 and 170 mir- loci are excluded from the Genome Browser. It is perhaps a temporary hiccup that may get resolved.
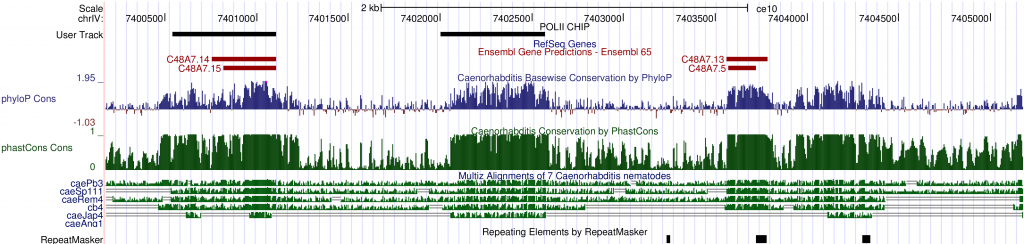
Once a region is selected, the Genome Browser can display a wide selection of sequences and features as tracks. The tracks are easily customizable. One can add or remove specific tracks and change the order or looks of tracks to make them easier to browse. Several tracks based on multiple genomic sequences are grouped under the Comparative Genomics section. There is a multiple sequence alignment track (Multiz) and three conservation tracks (phyloP, phastCons and conserved elements) to graphically display the results of conservation analyses. Scanning the genome with these tracks turned on, one can quite easily spot areas that show significant cross-species conservation, possibly implicating functional domains. For example, on chromosome IV, between C48A7.14 and C48A7.13, there is a 530-bp region (chrIV:7402060-7402590) that is highly conserved between C. elegansand five other species and yet there is no existing annotation. It could be an extension of a gene model nearby or a new gene.
One important extension of the Genome Browser is the ability to add custom tracks. Custom tracks allow users to incorporate their own or third party data into the browser so that imported sequence annotations can be inspected in the context of existing ones. For example, a large collection of diverse, sequence-based experimental data have been published by the ModEncode project. There are experiments assessing every part of the genome for its involvement in expression regulation under diverse biological conditions. ModEncode data sets may be downloaded as sequence features in a compatible format and included as custom tracks in the Genome Browser. As an example, I added a track for modencode_174 (labeled as “User Track: POLII CHIP”), which marks genomic regions that have been identified as potential binding sites for RNA polymerase II. There is substantial overlap between a potential Pol II binding site and inter-species conserved bases intervening C48A7.14 and C48A7.13. This observation further supports the possibility that this 530-bp stretch of sequence is functionally important, and may be expressed.
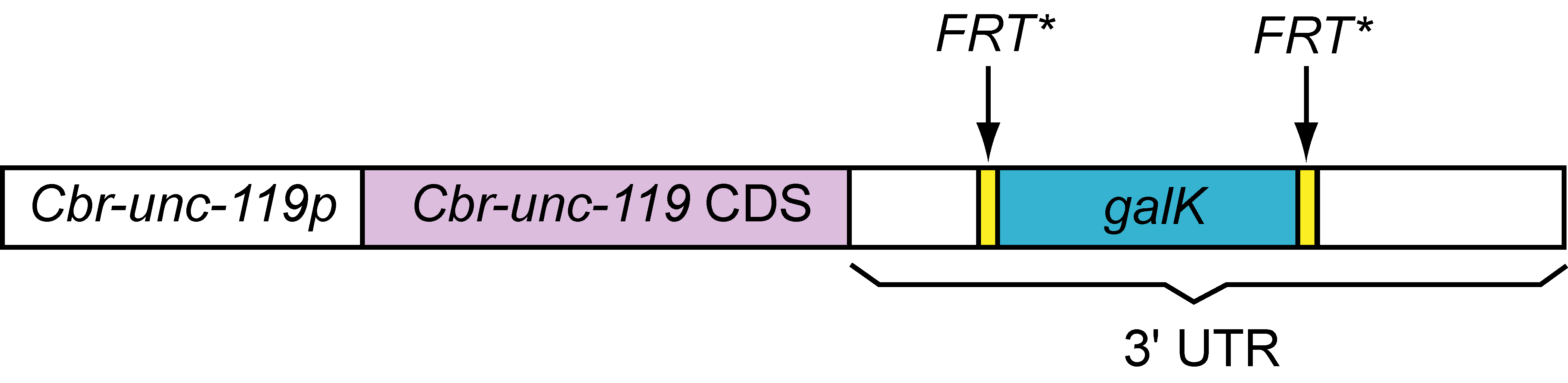
A universal Cbr-unc-119::FRT*::galk::FRT* template for recombineering and biolistics
 Table 1: Primers for amplifying the pPK719 cassette; pCC1FOS homology arms are shown in brackets. Note that the pCC1Fos homology sequence in PK951 is also present in the pBSKSII(+) vector backbone.
Table 1: Primers for amplifying the pPK719 cassette; pCC1FOS homology arms are shown in brackets. Note that the pCC1Fos homology sequence in PK951 is also present in the pBSKSII(+) vector backbone.
The 3.6 kb Cbr-unc-119::FRT*::galk::FRT* cassette, which is PCR amplified from pPK719 using the PK951/PK952 primer pair (Table 1), can be targeted by recombineering to insert between nts 281:282 in the pCC1FOS fosmid vector backbone. In other words, this cassette can be readily inserted in all fosmids contained in the C. elegans library developed by Don Moerman and colleagues. We have tested the construct in fosmid recombineering and have shown that it rescues unc-119(ed3) mutants. The clone will be made available at http://www.Addgene.com.
Figures
References
Maduro M, and Pilgrim D. (1996). Conservation of function and expression of unc-119 from two Caenorhabditis species despite divergence of non-coding DNA. Gene 183, 77-85. 
Praitis V, Casey E, Collar D, and Austin J. (2001). Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217-26. 
Tursun B, Cochella L, Carrera I, and Hobert O. (2009). A toolkit and robust pipeline for generation of fosmid-based reporter genes in C. elegans. PLoS One 4, e4625. 
A protocol for constructing and characterizing transgenic nematodes expressing stress-tolerant gene TPS1 of yeast in dauer larva stage
We are interested in constructing transgenic nematodes expressing the stress-tolerance gene TPS1 from yeast (Vellai et al., 1999), which is expressed in the dauer juvenile stage both in C. elegans and Heterorhabditis bacteriophora. In C. elegans experiments we used dauer constitutive mutants DR133 and DR136, homozygous for daf-7 and daf-2 respectively, and for morphological markers unc-32 (III) and dpy-1 (III). In H. bacteriophora experiments we used a 19th generation inbred line of H. bacteriophora strain TT01. C. elegans was grown on NGM, while H. bacteriophora was grown on ENGM (Fodor et al., The WBG 18:2). The TPS1 gene was isolated from S. cerevisiae by Z. Bánfalvi and ligated into the nematode vector (made by A. Fire) and cloned to the heat-inducible promoter hsp16-2 promoter, as described before (Vellai et al., 1999). Transformation was carried out by “bombarding.” In Hungary we have been using the Gene Booster equipment patented by Dr. B. Jenes in the Agro-Biotechnology Center, Gödöllő, Hungary. Synchronous L4 (J4) and dauer (IJ) populations were bombarded with different doses on bacterium-free NGM plates. The worms were washed off and transferred to fresh, bacterium seeded NGM (ENGM) plates. The NGM and ENGM plates were incubated in 18 and 25 oC, respectively. Bombarded generations considered to be P0. For C. elegans the next generation was allowed to grow and propagate on OP50-seeded NGM plates. Plates were transferred to 25 oC for 48h. The nematodes were washed off the plates and dauer larvae were selected in SDS. After 5X washing with sterile tap water they were heat shocked at 33 oC for 4h. The F1 progeny of the bombarded worms were transferred to 2.4 M NaCl for 4h. 100% of the control worms died during this time. The progeny of the bombarded worms were washed 3X and distributed to 40 NGM plates seeded with OP50 bacteria. Some survived. They are considered as potentially transformed nematodes. One individual hermaphrodite was picked up from each plate to initiate pure lines for further study. For H. bacteriophora each ENGM plate, like an island, was put in a large glass plate of sterile tap water. The lid of the ENGM plate was removed, but the large plate covered. The IJ were moving from the plates to the water. They were collected daily. They were handled similarly to C. elegans except for using 0.1 w/v hyamin instead of SDS. The Fn progeny of the bombarded worms were transferred to 2.4 M NaCl for 4h. 100% of the control died during this time. The animals were washed 3X with sterile tap water and distributed into 40 ENGM plates seeded with TT01 bacteria. Some survived. They were considered as potentially transformed nematodes. One individual hermaphrodite was picked up from each plate to initiate pure lines for further study. We found no significant difference in the rate of survival between the progeny of treated L4 (J4) and dauers.
References
Vellai T, Molnár A, Lakatos L, Bánfalvi T, Fodor A, and Sáringer G. (1999). Transgenic nematodes carrying a cloned stress resistance gene from yeast. In Survival of entomopathogenic nematodes. Glazer I, Richardson P, Boemare N, and Coudert F, eds. (Luxembourg, European Commission Publications), pp. 105-119. 
Unnatural amino-acid tagging and read-through of nonsense stop codons in multicopy transgene arrays
Incorporation of unnatural amino-acids is a powerful and versatile new tool for modifying and selectively tagging proteins of interest. The technique has been shown to be applicable to C. elegans by Greiss and Chen (2011), utilizing a transgene expressing an amber-suppressing tRNA together with a cognate pyrrolysl-tRNA synthetase from Methanosarcina.
We wished to apply this technique to investigate srf-5, a gene affecting surface antigenicity and bacterial infection susceptibility, which we have found to encode a predicted small secreted protein. The reference srf-5 allele, ct115, causes an early Trp-to-opal nonsense mutation and behaves as a genetic null. We reasoned that it should be possible to rescue the mutant phenotype by introducing a transgene carrying a Trp-to-amber mutation at the same position, and crossing this transgene onto an amber-suppressing genetic background such as sup-5. Crossing the same transgene onto the unnatural-amino-acid-inserting background developed by Greiss and Chen (2011), and providing a suitable unnatural amino-acid substrate, should then result in synthesis of SRF-5* (that is, SRF-5 with an unnatural amino acid instead of Trp). If SRF-5* is as functional as wildtype SRF-5, then it should also rescue the srf-5 mutant.
Surprisingly, we found that the srf-5(amber) transgene was able to rescue the srf-5 mutant in the absence of any amber suppression, when present in a multicopy extrachromosomal array. The same effect was seen when the original opal nonsense mutant, ct115, was introduced as a multicopy extrachromosomal array. We hypothesized that transcription from the multiple copies leads to elevated srf-5 mRNA levels, and consequently to some read-through of either stop codon. We tested this interpretation by crossing srf-5(ct115) with smg-2(e2008), a mutant defective in NMD (Nonsense Mediated Decay), and found that smg-2 also partly suppressed the Srf-5 phenotype. The ct115 opal (UGA) codon therefore appears to be a leaky stop. It may be unusually leaky, or else SRF-5 may be required only in very small amounts. An alternative explanation, that the amber fragment encoded by srf-5(ct115) has some functional activity, seems unlikely because this fragment is small (<35 amino-acid residues).
Using amber transgenes for unnatural amino-acid incorporation should work for other genes, and may still work for srf-5 itself. However, the read-through effect that we report here may sometimes complicate matters.
Artifacts in L1 starvation assay
Survival of L1 arrested larvae in the absence of food is a common experiment in C elegans lifespan studies. A typical protocol involves resuspension of freshly bleached worm eggs in sterile inorganic buffer. Newly hatched worms arrest due to lack of food and their starvation survival in the following days is monitored by taking aliquots and counting live worms. It is tempting to think that such well defined and simple experimental conditions leave little room for variability. In reality, the outcome of this experiment is sensitive to a number of often neglected factors ranging from worm maintenance history dating several generations back to small temperature fluctuations during starvation. Recently, we found additional sources of artifacts in L1 survival assays.
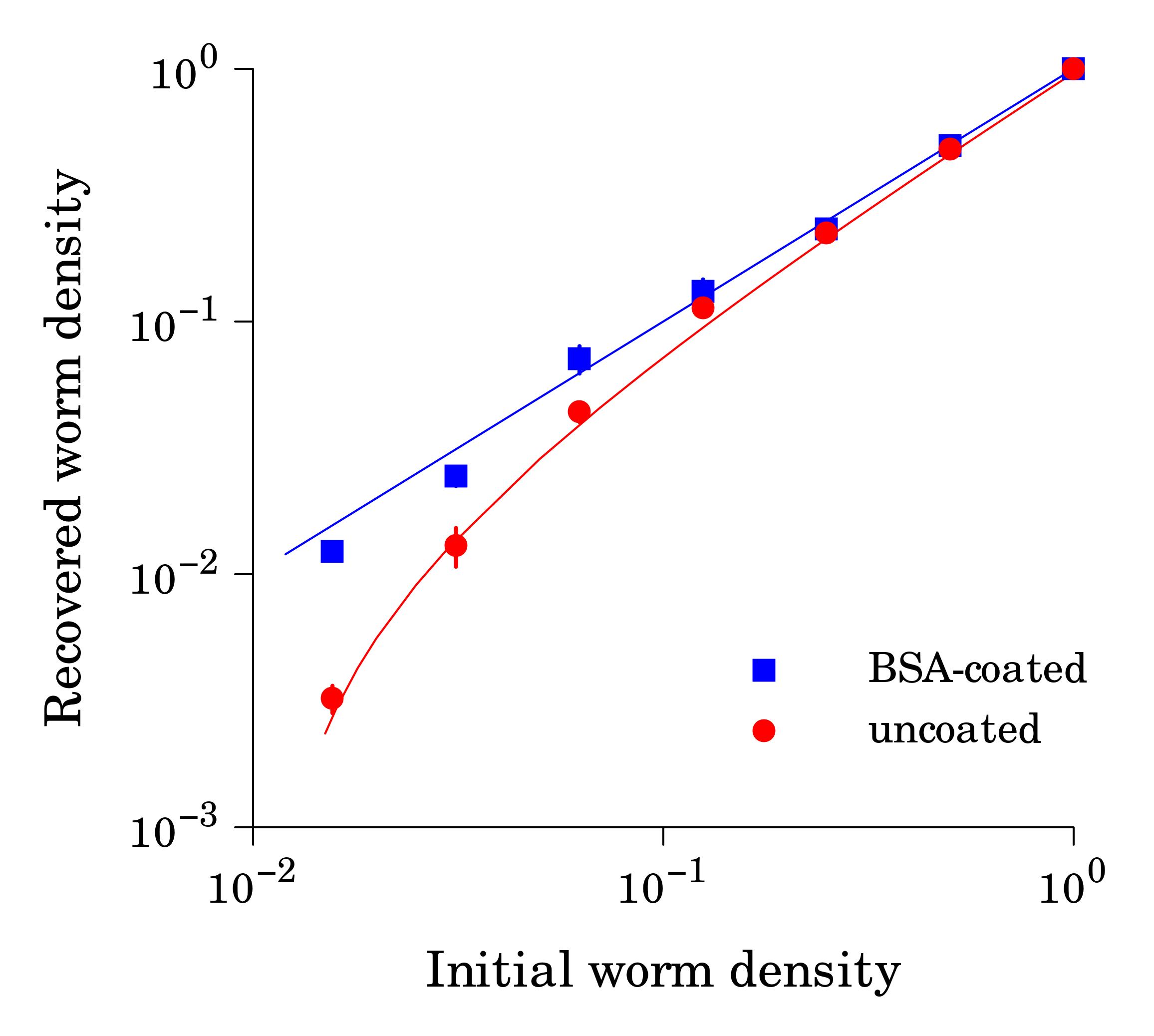
When starved L1 larvae are incubated in M9 buffer in plastic tubes (e.g. polypropylene 15-ml tubes), worms tend to adhere to hydrophobic plastic walls, which decreases apparent worm counts in suspension. This effect becomes particularly noticeable at low worm densities (< 3 worms/μl, Fig. 1) We found that precoating tubes with bovine serum albumin (1% aqueous solution for an hour followed by 3 water rinses) renders plastic hydrophilic and greatly reduces the sticking problem. BSA solution and water for rinses should be sterile.
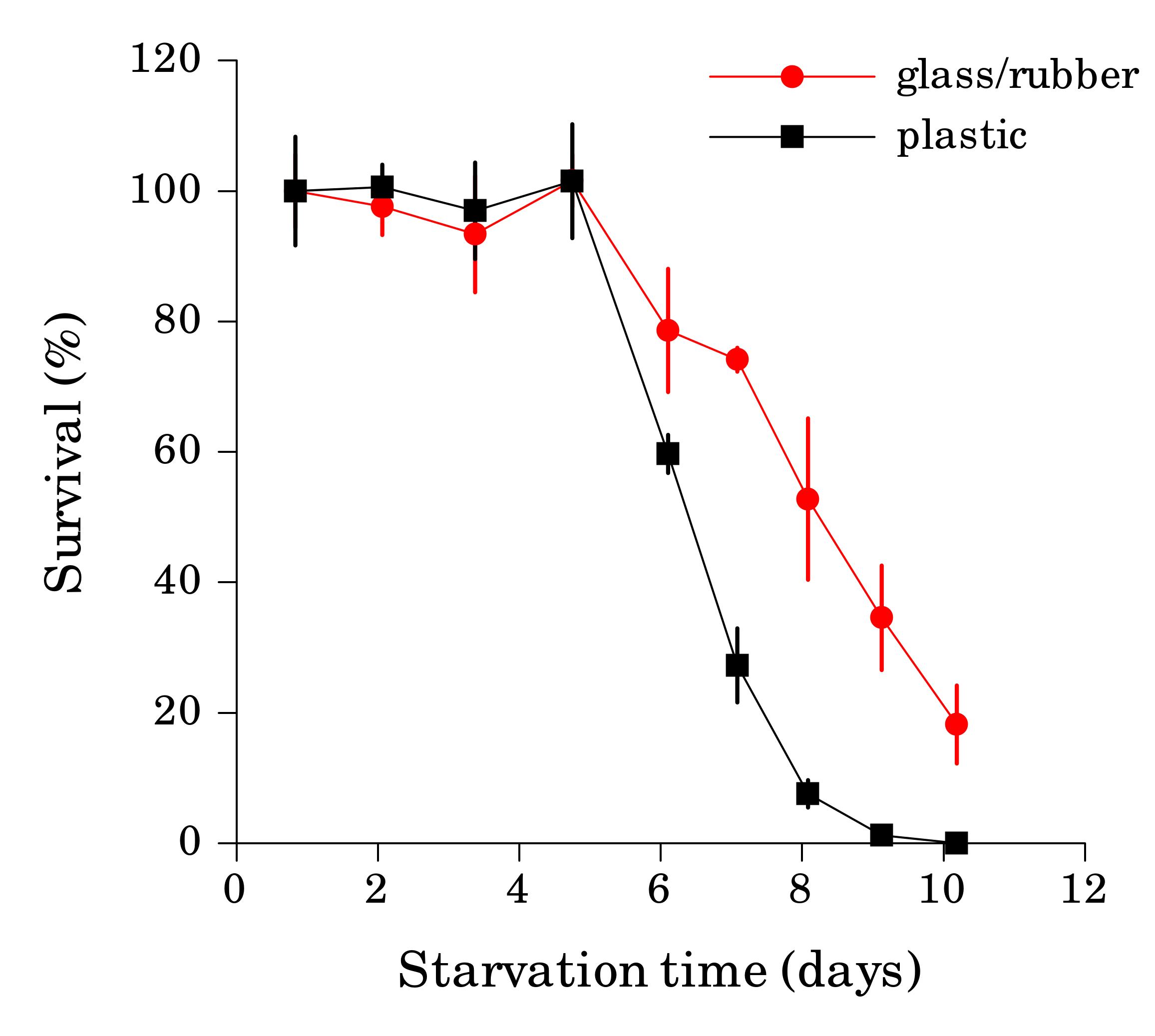
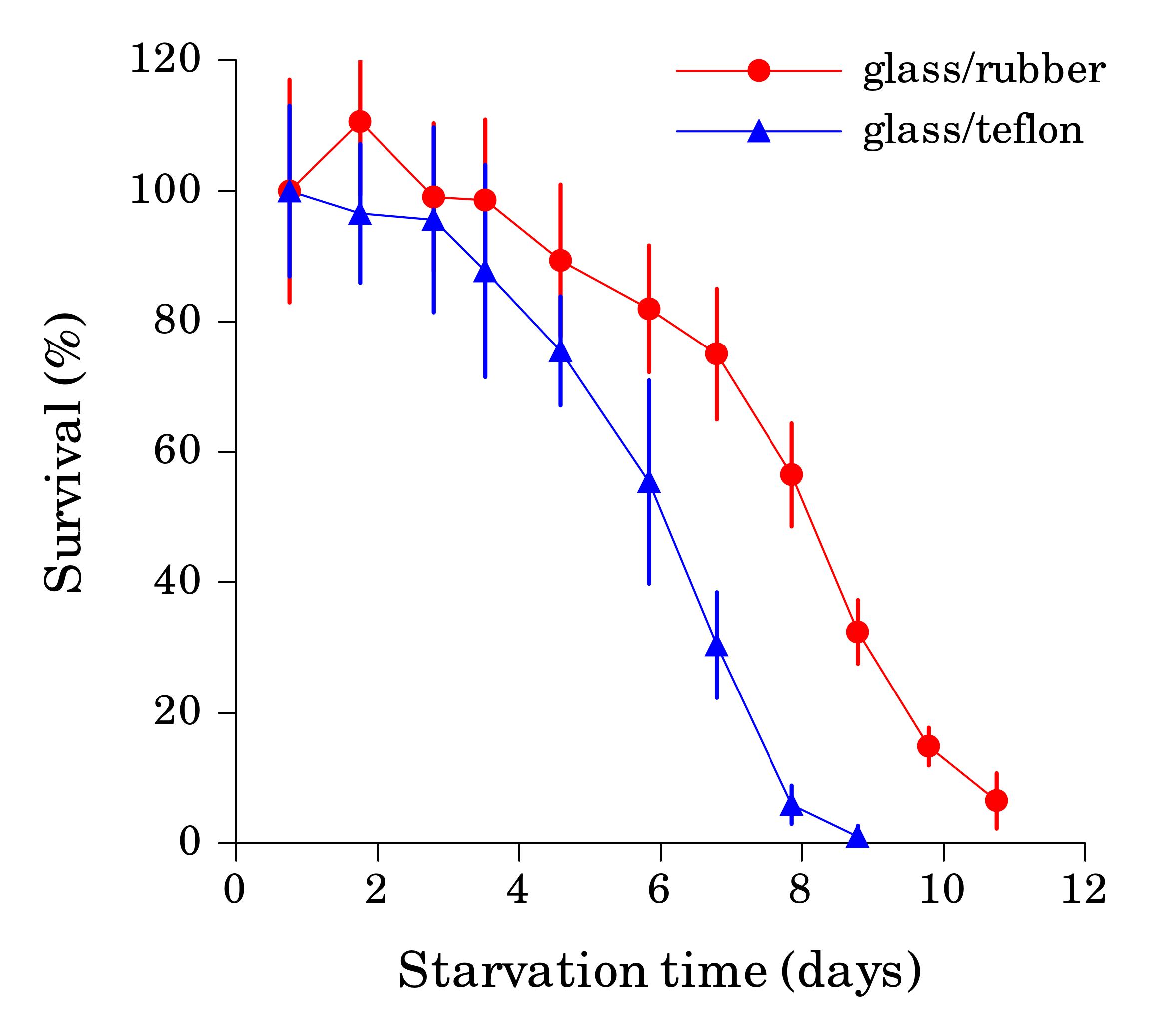
When the starvation experiment is performed in glass tubes or vials, worm sticking is not a problem, since clean glass is hydrophilic. The danger in this case comes from the cap, since seemingly inert material lining the cap can significantly affect survival rate if it comes in contact with liquid. We found that L1 worms starved in glass tubes with rubber-lined caps reproducibly survived starvation longer then worms in plastic tubes of the same volume (Fig. 2) or glass tubes with teflon-lined caps (Fig. 3). We detected several compounds in water from tubes with rubber-lined caps and identified one of them as 2-mercaptobenzothiazole (2-MBT), which was present at ca. 0.5 μM. 2-MBT, used as an accelerator in the vulcanization of rubber, is commonly seen in rubber leachates (Reepmeyer and Juhl, 1983). We speculate that longer survival is due to a hormetic effect of one or several chemicals leaching from rubber, which would be toxic at higher concentrations. However, low concentrations of synthetic 2-MBT alone were not sufficient to reproducibly extend L1 starvation survival (at 3 μM and above 2-MBT is toxic to worms). Other components of the rubber leachate may be necessary for the effect.
These results demonstrate artifacts that can both decrease and increase apparent survival rates in L1 starvation.
Figures