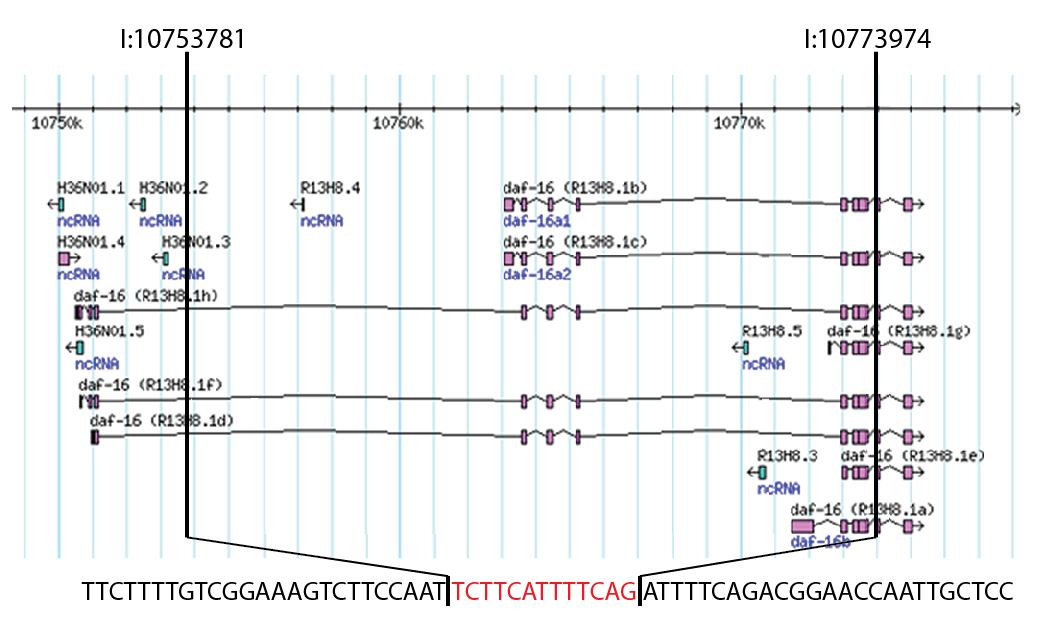
To map the endpoints of the deletion region, we utilized nested PCR and designed a series of oligos based on the described breakpoints of the deletion region (Ogg et al., 1997). We thought that the 5’ endpoint would occur within 2 kb of the start codon for the R13H8.16b and R13H8.1c isoforms, which are also commonly referred to as daf-16a1 and daf-16a2. However, these oligos failed to amplify genomic DNA from the mgDf50 mutant, so we tested oligos that hybridize further upstream. With the oligos F (5’-AAATATGGGAATATCATATCGAAAA-3’) and R (5’-AATTCCAGGCAGTGGAGATGA-3’), we successfully obtained a ~2,000 base pair PCR product which we subcloned and sequenced using vector derived oligos. Sequence alignment demonstrated that the deletion spans from position 10753781 to 10773974 on chromosome I and also includes a 13 base pair insertion. As shown in Figure 1, this 20,193 base pair deletion starts within a large intron present in the daf-16h, d, and f isoforms, spans the start codons for the daf-16a isoforms, and ends at the beginning of exon 5 for the R13H8.1a isoform, which is often called daf-16b. Importantly, the exons encoding the DNA binding domain are included in the deletion.
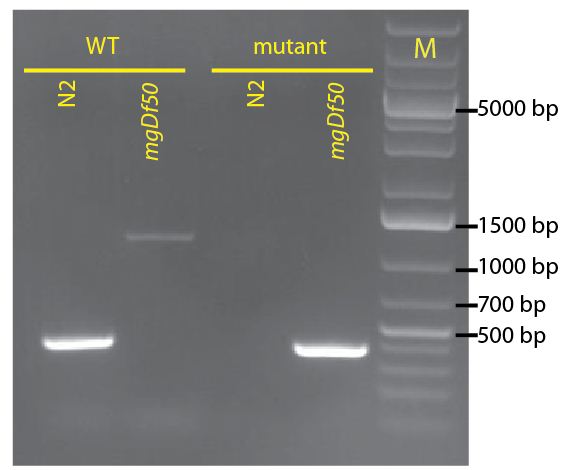
To identify the daf-16(mgDf50) allele, we use the oligos: mgDf50-F (5’-TGCCCTCTCTCTGTTTCTCC-3’) and mgDf50-R (5’-GATTCTGAGCACACGATCCA-3’), which flank the deletion region in the mgDf50 allele and amplify a 411 base pair band; and daf-16WT-F (5’-TCTACGGTTGGAACGGATTC-3’) and daf-16WT-R (5’-CCCCATGGTGTTCTCACTCT-3’), which hybridize inside the daf-16(mgDf50) deletion region and amplify a 389 base pair band from the wild type allele. Further, the WT oligos hybridize inside an intron in the daf-16 sequence, so they can be used even in the presence of cDNA based transgenes, such as those recently described by Kwon et. al. and now available at CGC (Kwon et al., 2010). Figure 2 shows the genotyping of a homozygous daf-16(mgDf50) and wild-type strain with these oligos.
Figures
References
Lin K, Dorman JB, Rodan A, Kenyon C. (1997) daf-16: An HNF-3/forkhead family member that can function to double the life-span of Caenorhabditis elegans. Science 278, 1319-1322. 
Hsin H, Kenyon C. (1999) Signals from the reproductive system regulate the lifespan of C. elegans. Nature 399, 362-366. 
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R. (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366, 461-464. 
Kwon ES, Narasimhan SD, Yen K, Tissenbaum HA. (2010) A new DAF-16 isoform regulates longevity. Nature 466, 498-502. 
Ogg S, Paradis S, Gottlieb S, Patterson GI, Lee L, Tissenbaum HA, Ruvkun, G. (1997) The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389, 994-999. 
Riddle DL, Swanson MM, Albert PS. (1981) Interacting genes in nematode dauer larva formation. Nature 290, 668-671. 
Articles submitted to the Worm Breeder's Gazette should not be cited in bibliographies. Material contained here should be treated as personal communication and cited as such only with the consent of the author.
Leave a Reply
You must be logged in to post a comment.