In common with many labs, in the past we routinely used DAVID to look for over-representation of functional classes in lists of genes. Unfortunately, the data in DAVID hasn’t been up-dated since September 2009. As an alternative, there’s a stand-alone version of DAVID called EASE. This allow labs to use their own lists. A post to the forum last year asking whether anyone maintained up-to-date EASE lists went unanswered, so we decided to make our own. That’s when we realized that there are three challenges. One is simply having a user-friendly interface to enter any new gene list, with the appropriate citation information. The second is converting any gene list culled from the literature into a coherent form. This is made more difficult by the fact that one still finds gene lists in published papers that (i) contain an apparently random mix of different types of gene identifiers and (ii) don’t say which WormBase WS version was used. The third is that with changes in predicted gene structure (genes being split, killed, fused, etc), lists become increasingly inaccurate with time.
We therefore made a first tool, the WormBase Converter. It can automatically detect different IDs (Transcript Name, Gene Name, WormBase ID, etc.) and convert them into the type of ID you need. It also allows the conversion between WormBase releases. You can convert a Gene ID from a specific release (e.g. WS170) into another release (e.g. WS220 for a conversion, or WS160 for a reverse-conversion).
As WormBase Converter was originally made for use with EASE, we didn’t design it to give a correspondence table, but are currently working on adding this as an option, so that there will be the possibility of having an output table listing in one column the input genes and in an adjacent column the list of genes in the required output format. There are stand-alone WormBase Converter versions for all common platforms (Unix, MacOS, Windows) available on Sourceforge.org. As keeping it up to date does take some effort, due to the need to correct the occasional Wormbase annotation anomaly, we also offer a client version that sends requests to the WormBase Converter server that we maintain at the CIML. We’d be delighted if anyone else would like to host this service!
We also made EASE Manager. This facilitates the entry and management of gene lists into EASE, but more importantly, interfaces with the WormBase Converter so that the genes that make up any list are kept up to date. Installation instructions and full documentation can be obtained from our website. We welcome feedback on these tools, which we also described in a paper earlier this year (Engelmann et al., 2011).
Figures
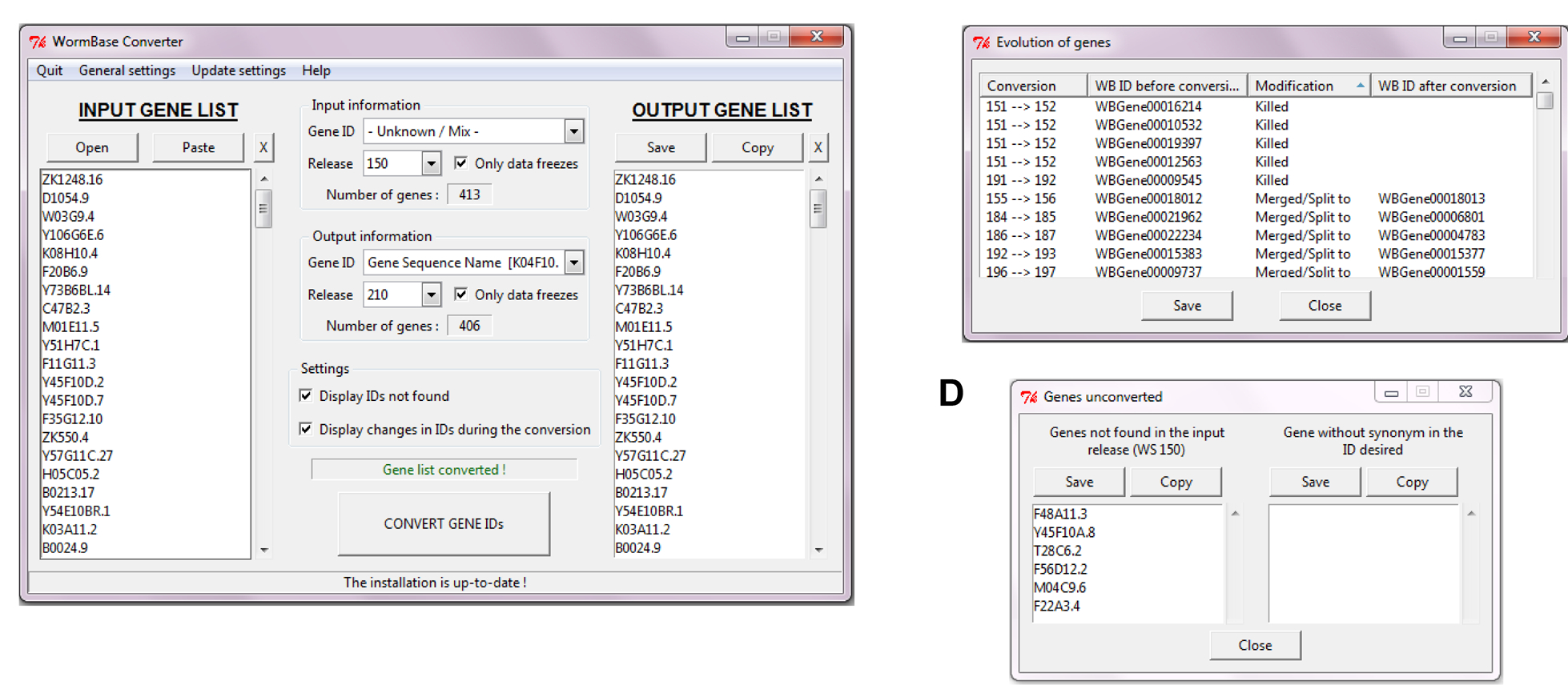
References
Engelmann I, Griffon A, Tichit L, Montañana-Sanchis F, Wang G, Reinke V, Waterston RH, Hillier LW and Ewbank JJ. (2011). A comprehensive analysis of gene expression changes provoked by bacterial and fungal infection in C. elegans. PLoS One 6, e19055. 
Articles submitted to the Worm Breeder's Gazette should not be cited in bibliographies. Material contained here should be treated as personal communication and cited as such only with the consent of the author.
Leave a Reply
You must be logged in to post a comment.