We previously published methods to allow worm researchers to
(i) identify potentially useful Mos1 insertions from the NEMAGENETAG collection (MosLocator)
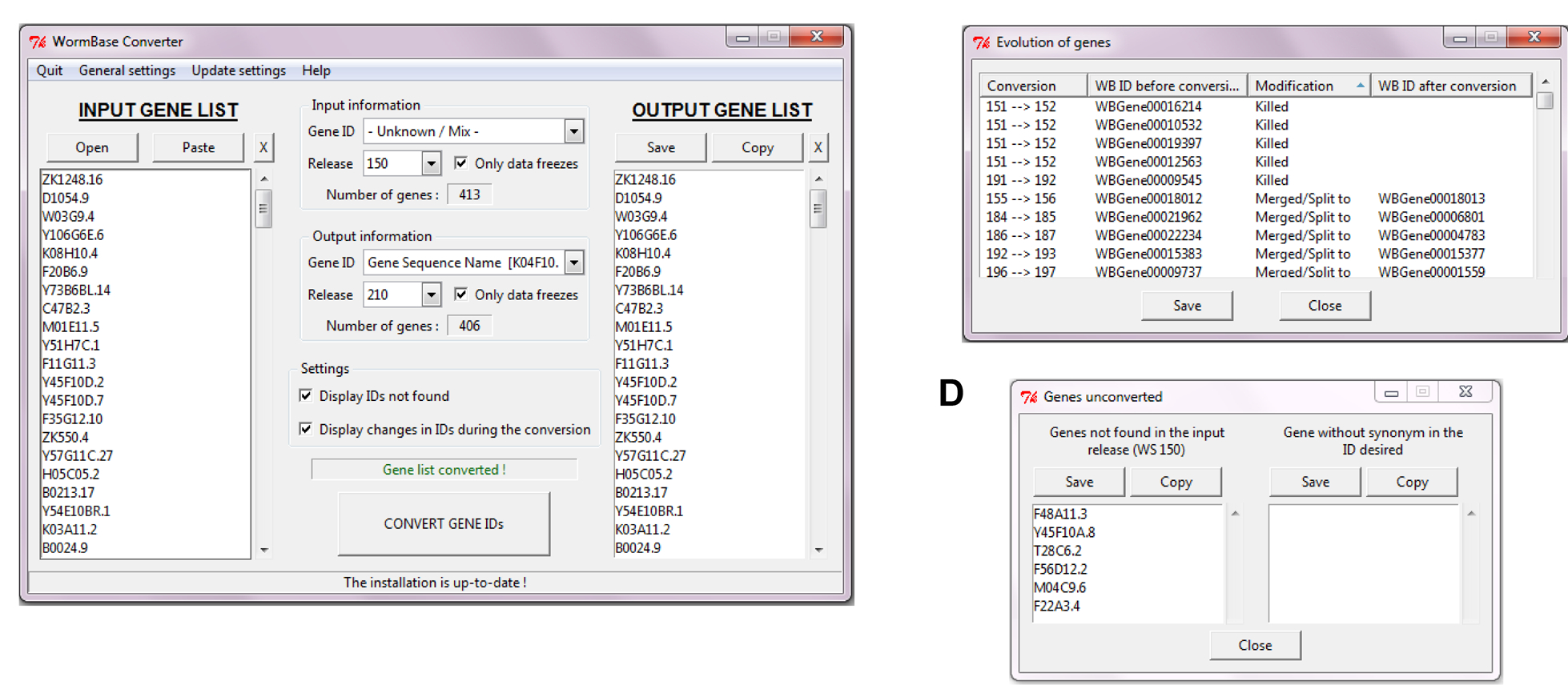
(ii) convert and/or update lists of genes (WormBase Converter; see Griffon and Ewbank, WBG 18 #4)
We’ve now added 2 more tools specifically designed for worm labs, with the aim of making life easier:
– ICeE, a worm-centered electronic notebook that facilitates storage of data and metadata. It comes pre-loaded with the CGC list of strains and the RNAi clones from the main collections.
– Clone Mapper, which does three things:
1. Map your clones: Automatically determines whether a clone taken from the Ahringer or Vidal RNAi library is what it is supposed to be, by mapping your clone sequences to an in-silico RNAi clone library.
2. Find Targets: Predicts gene targets for RNAi clones, on the basis of matching n-mer sequences, overcoming some of the limitations of current Wormbase predictions.
3. Retrieve Sequences: Allows single or multiple clone and transcript sequences to be retrieved.
Many people contributed to the elaboration of these tools, and we benefited from invaluable input from numerous members of the WormBase staff.
All these tools are freely available via our website. Feedback is welcome.
If you use any of them, please do cite the relevant publications:
MosLocator: Vallin et al., 2012 PMID: 22347378
Wormbase Converter: Engelmann et al., 2011 PMID: 21602919
ICeE: Montanana et al., 2014 http://dx.doi.org/10.4161/worm.32160
Clone Mapper: Thakur et al., 2014 PMID: 25187039
References
Engelmann I, Griffon A, Tichit L, Montañana-Sanchis F, Wang G, Reinke V, Waterston RH, Hillier LW, Ewbank JJ. (2011). A comprehensive analysis of gene expression changes provoked by bacterial and fungal infection in C. elegans. PLoS One 6, e19055. 
Frédéric Montañana, Renaud A Julien, Philippe Vaglio, Lisa R Matthews, Laurent Tichit, Jonathan J Ewbank (2014). ICeE: an interface for C. elegans experiments. Worm 3, e32160. http://dx.doi.org/10.4161/worm.32160 BioRXiv
Thakur N, Pujol N, Tichit L, Ewbank JJ. (2014). Clone Mapper: an online suite of tools for RNAi experiments in Caenorhabditis elegans. G3 (Bethesda) pii: g3.114.013052.doi: 10.1534/g3.114.013052. 
Vallin E, et al., (2012). A genome-wide collection of Mos1 transposon insertion mutants for the C. elegans research community. PLoS One 7, e30482.