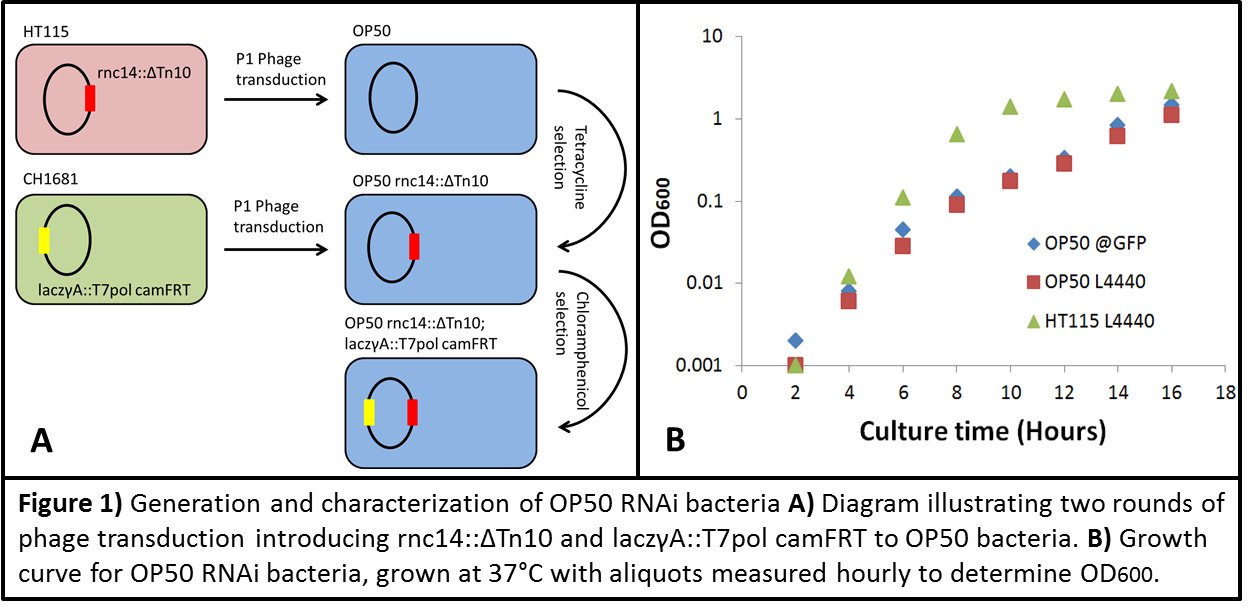
In the majority of research laboratories using C. elegans as a model, we sustain our animals on the B-type E. coli strain OP50 (Brenner, 1974). This strain has provided us with a historically key platform for defining genetic pathways. Concurrently, the technique of double stranded RNA (dsRNA) feeding to incite RNA interference (RNAi) has become a key technique in many laboratories. Two major resources, the Ahringer and Vidal libraries house thousands of E. coli clones with unique plasmids expressing dsRNA corresponding to individual C. elegans genes (Kamath and Ahringer, 2003; Rual et al., 2004). While numerous experimental insights have been facilitated using these resources, both libraries are housed in the HT115 E. coli K12 strain. Several recent studies (Brooks et al., 2009; Soukas et al., 2009; Pang and Curran, 2014) have revealed the importance of the bacterial environment in eliciting certain C. elegans phenotypes. With the realization that gene-environment interactions may play a key role in genetic regulation, we have generated an OP50 strain capable of dependably housing plasmid vectors expressing dsRNA. Furthermore, when C. elegans are fed this OP50 strain, they activate a robust RNAi response against the target gene of interest. We anticipate that this resource will allow the further characterization of gene-environment specific relationships in C. elegans. To generate an RNAi competent OP50 strain, two main steps were required. We started by replacing the wild-type (WT) OP50 allele of RNAIII RNase (rnc) with the same deletion allele found in the E. coli K12 HT115 strain. We then introduced a genomically encoded IPTG-inducible T7 RNA polymerase to drive double stranded RNA production from L4440 plasmids. To do this, we prepared P1 phage lysates from strains HT115 (rnc14::DTn10) and CH1681 (laczgA::T7pol camFRT) (Figure 1A). We initially transduced OP50 overnight (O/N) culture with the HT115 (rnc14:: DTn10) lysate and selected for tetracycline (tet) resistance. After repetitive rounds of selection, we inoculated individual colonies into LB + tet and confirmed the presence of the rnc14::DTn10 allele by PCR. Following this, we transduced OP50 (rnc14::DTn10) O/N culture with CH1681 (laczgA::T7pol camFRT) P1 lysate and used multiple rounds of Chloramphenicol (cam) positive selection to purify our background. Individual colonies were then picked, and PCR was used to confirm the presence of rnc14:: DTn10 and laczgA::T7pol camFRT.
Grown overnight, OP50 replicative timing tends to be slightly delayed in comparison to its HT115 counterpart. However, the desired ODs can be easily reached which facilitate the induction of an RNAi response (Figure 1B).
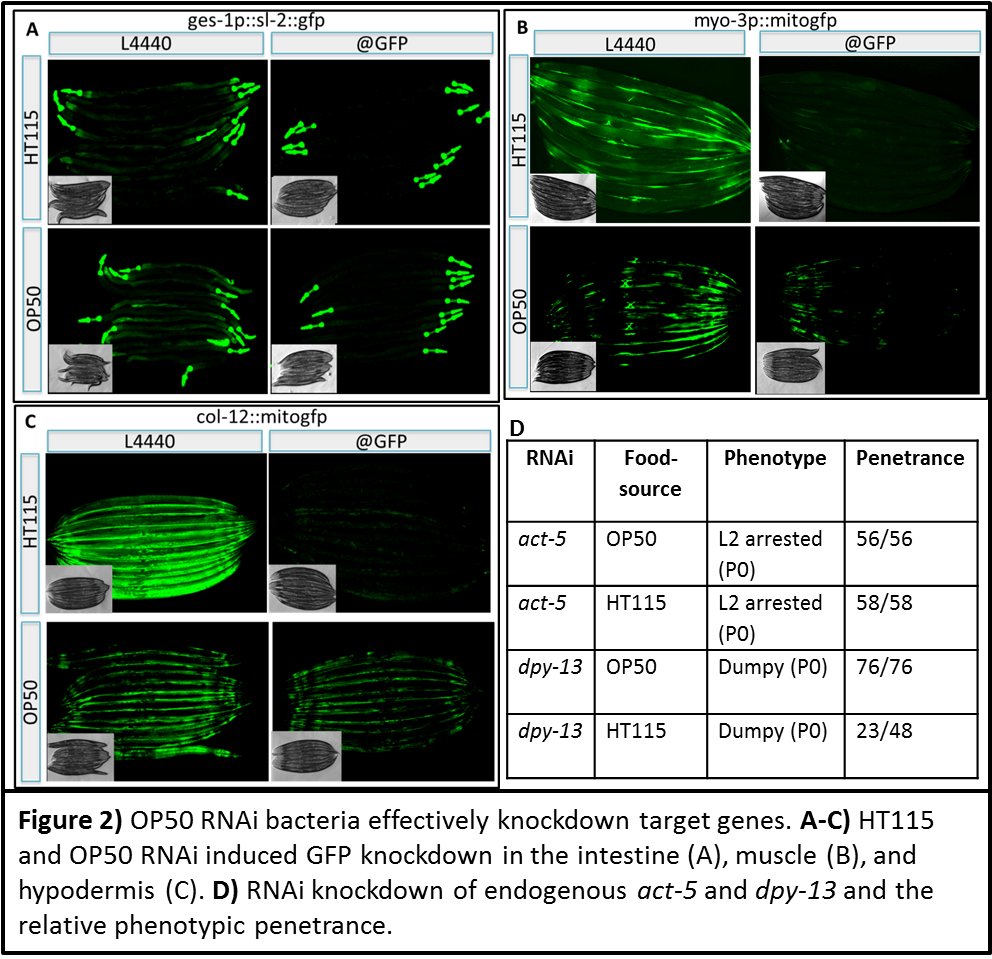
Using Pges-1::sl2::gfp as our reporter C. elegans strain, we determined that after 16h of culture, OP50 RNAi competent strains housing the L4440 plasmid targeting GFP are capable of inducing significant knock-down of the target gene. This knockdown was then repeated in both the muscles and hypodermis (Figure 2A-C).
Next we generated OP50 RNAi clones targeting two endogenous C. elegans genes: act-5 and dpy-13. In the P0 generation, OP50 derived dsRNA knockdown of act-5 is as sufficient as its contemporary HT115 strain in arresting worm development. RNAi knockdown of dpy-13 in using the OP50 background exerts an even stronger effect in the reduction of worm size than the comparable HT115 knockdown experiment (Figure 2D).
Thus we have constructed an OP50 strain which can robustly initiate RNAi reactions comparable to those generated by HT115. The knockdowns generated can be in multiple different tissues, and can target various genes. Utilizing this method we plan to further elucidate gene-environment interactions. This resource is available to the research groups requesting it.
References
Brenner S. (1974). The genetics of Caenorhabditis elegans. Genetics 77, 71-94. 
Kamath RS, Ahringer J. (2003). Genome-wide RNAi screening in Caenorhabditis elegans. Methods 30, 313-321. 
Rual JF, Ceron J, Koreth J, Hao T, Nicot AS, et al. (2004). Toward improving Caenorhabditis elegans phenome mapping with an ORFeome-based RNAi library. Genome Res. 14, 2162–2168. 
Brooks KK, Liang B, Watts JL. (2009). The influence of bacterial diet on fat storage in C. elegans. PLoS One, 4:e7545. 
Soukas AA, Kane EA, Carr CE, Melo JA, Ruvkun G. (2009). Rictor/TORC2 regulates fat metabolism, feeding, growth, and life span in Caenorhabditis elegans. Genes Dev. 23, 496–511. 
Pang S, Curran SP. (2014). Adaptive capacity to bacterial diet modulates aging in C. elegans. Cell Metab. 19, 221-31. 
Articles submitted to the Worm Breeder's Gazette should not be cited in bibliographies. Material contained here should be treated as personal communication and cited as such only with the consent of the author.
Leave a Reply
You must be logged in to post a comment.