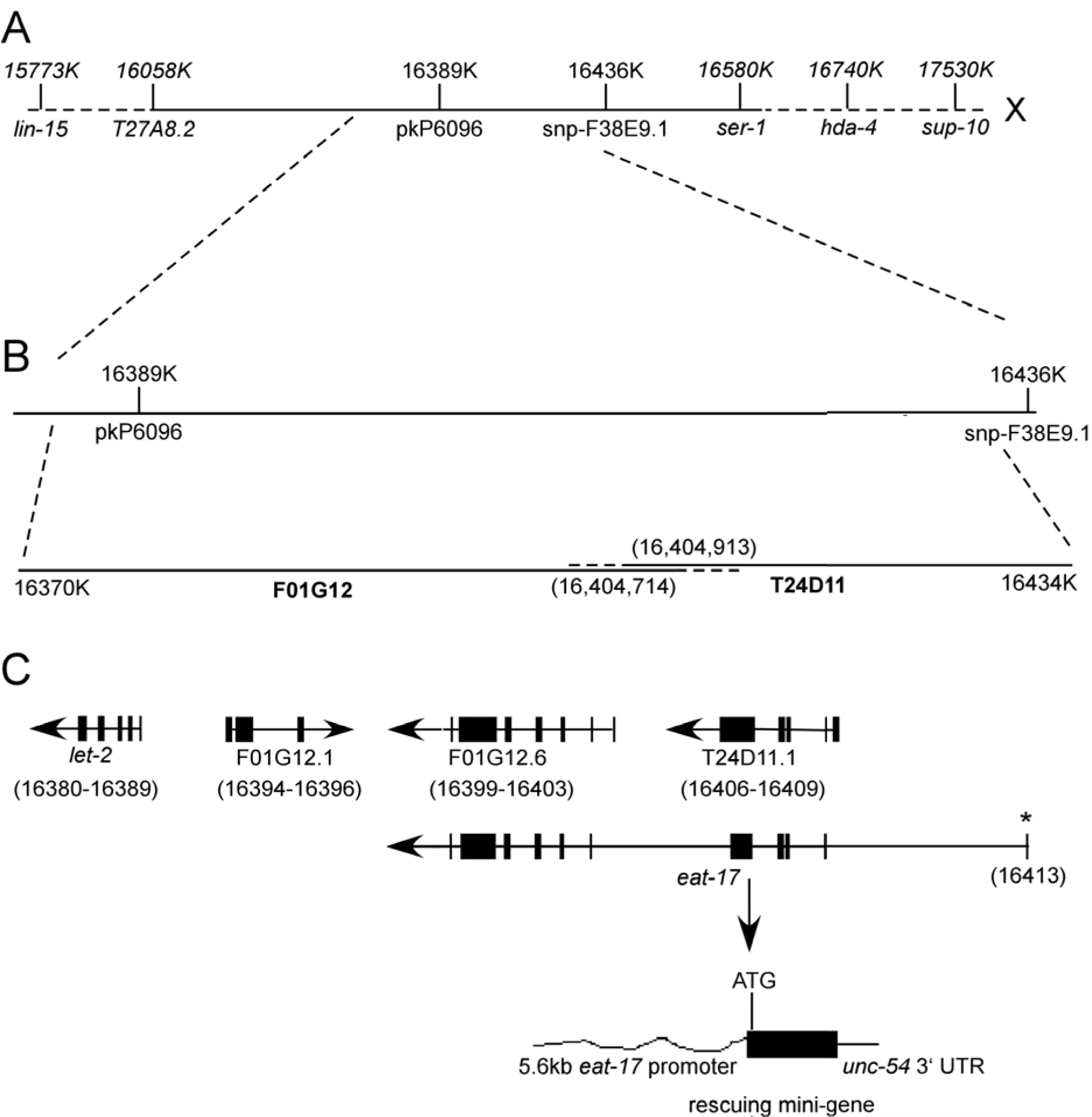
eat-17 was originally mapped to the right arm of Chromosome X, between lin-15 and sup-10. We used a three point mapping strategy combined with SNP mapping to narrow this down to a region between pkP6096 and snp-F38E9 (Figure 1). Co-injection of cosmids T24D11 and F01G12 was both necessary and sufficient to rescue feeding defects in a subset of transgenic progeny. One predicted gene in this region—with homology to the human Evi5 gene—became a strong candidate. Three key findings were used to show that this was the eat-17 gene. First, amplification and injection of the cDNA corresponding to this region was able to rescue the eat-17 mutant phenotype. Second, RNAi against this region in rrf-3 mutants resulted in phenocopy of the eat-17 mutant phenotype. Finally, we found a C to T base pair transition at nucleotide 607 in exon 5, creating a stop codon at amino acid 203, potentially rendering the protein non-functional.
Based on homology searches, we found that EAT-17 belongs to a family of Rab GAPs. By RT-PCR, we identified two transcripts, differing by 9 nucleotides. Two predicted proteins, 825aa and 828aa, respectively, are encoded by these two transcripts (Figure 2). Each contains a predicted RabGAP domain at the N terminus and a series of coiled coil domains at the C terminus. We used RNAi studies to identify RAB-6.2 as EAT-17’s interacting partner. RNAi of both eat-17 and rab-6.2 leads to defects in grinder formation. Though the role of these proteins in grinder formation is not clear, we suspect that both play an important role in trafficking collagens, chitin, or other components of the grinder to the apical surfaces of the terminal bulb muscles, particular during molts. Defects in other chitin based components, including the sieve, may explain defects in food trapping.