The modENCODE ‘Integrative Analysis’ paper (Gerstein et al., 2010) described 304 unusual transcription factor binding site regions which they named ‘Highly Occupied Target (HOT) regions’. These short (~400 bp) regions bind to 15 or more transcription factors and are not enriched in transcription factor motifs when compared to the normal specific binding sites. The HOT regions are associated with genes which are highly expressed and which are more likely to be essential compared to other genes. The HOT regions are included in WormBase as the features ‘WBsf216780’ to ‘WBsf217083’, inclusive.
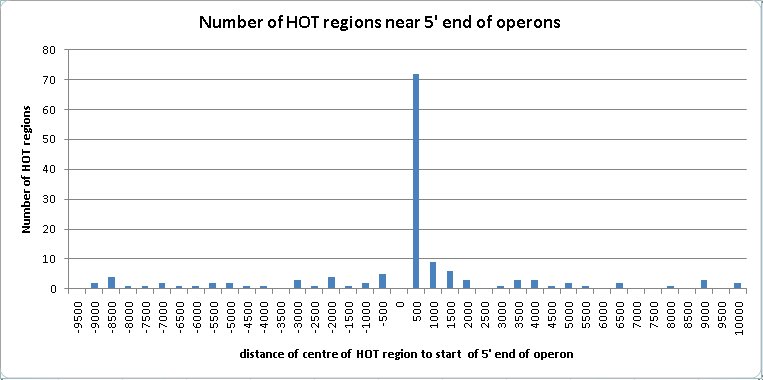
The HOT regions tend to be associated with the 5′ region of operons. This association was not commented upon by the authors of the modENCODE paper (Gerstein et al., 2010).
82 of the HOT regions are located within 2 Kb of the 5′ end of operons. 20 of these are within 2 Kb of two operons flanking them on both strands. Operons sometimes have genes at either end which have not been included in them for lack of evidence, or have genes added to them which should not be included. If the distance from the start of the operon is extended to 10 Kb to allow for the uncertainty in the start position of an operon, then the number of HOT regions located near the 5′ end of an operon increases to 143, and the number of these with two flanking operons increases to 40.
Assuming the null hypothesis that HOT regions should associate with the genes at the 5′ end of operons as frequently as with any other coding gene, the binomial distribution gives a significant result for the observed association (p-value < 2.2e-16).
HOT regions may be useful for locating operons which have not been curated so far. Several such potential operons have been seen during a cursory inspection of those HOT regions which are not near a known operon. HOT regions will not however be used by the WormBase curators as evidence for the existence of a nearby operon because HOT regions are just as likely to control genes with a property like being constitutively expressed (which could preferentially include operon genes) as they are to be specific markers for the 5′ end of operons.