There are about 20,000 protein-coding genes in C. elegans (WormBase WS238). We previously generated OrthoList, a compendium of C. elegans genes with human orthologs, by a meta-analysis of the results of four different orthology-prediction programs (Shaye and Greenwald, 2011.). Through compiling OrthoList, we found that 7,663 genes have predicted human orthologs, corresponding to about 38% of the worm genome. We also described how OrthoList addressed various disparities in, and limitations of, prior studies and validated it as a potential tool for streamlining RNAi screens.
We originally provided OrthoList as a spreadsheet with multiple tabs. We have now made a simple online tool that facilitates its use for determining worm-human orthology and generating personalized lists of genes of interest for RNAi screens or other purposes. The tool is available at http://www.greenwaldlab.org/ortholist.
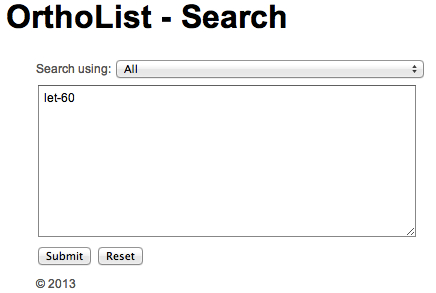
A search window (Figure 1) allows input of worm or human genes, one or many at a time. Worm genes should be introduced as WBgene IDs, common names or locus IDs to retrieve the predicted human ortholog(s). Human genes should be introduced as Ensembl ENSG IDs to retrieve the predicted C. elegans ortholog(s).
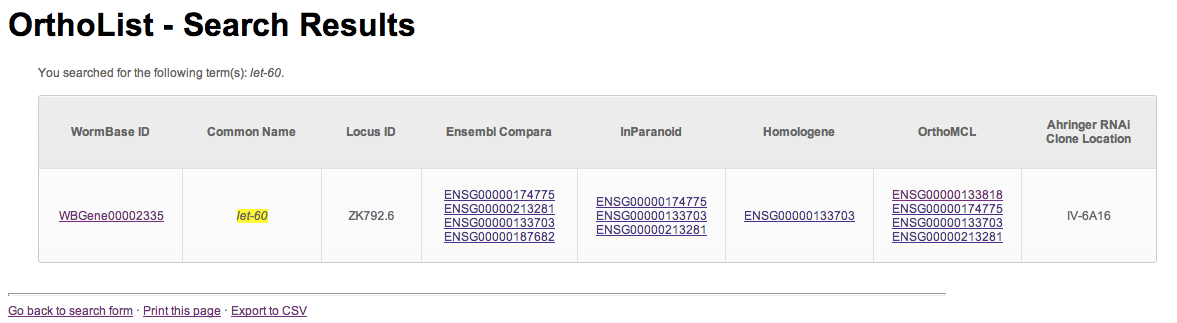
Results are returned as a table (Figure 2), in which the search term(s) are highlighted and orthologs are listed, along with which program(s) predicted the relationship. If a feeding RNAi clone is available, its location is also listed. Results can be exported to a CSV document, which can be read by text-editing and spreadsheet programs.