Worm Breeder's Gazette 9(3): 67
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
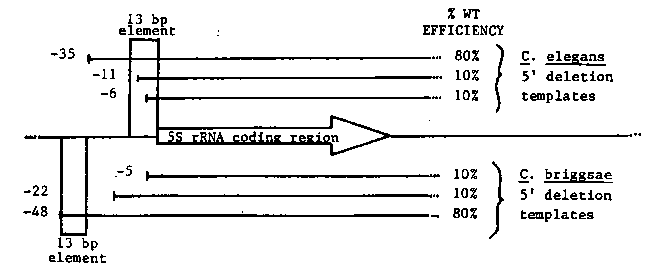
Cell free extracts of C. support RNA pol III specific transcription of circular plasmid templates carrying C. sequences encoding 5S or tRNAs. We are engaged in the identification of template sequences required for efficient 5S rRNA transcription in vitro. We have constructed a set of 5' and 3' deletion derivatives of a transcriptionally active C. DNA repeat. The efficiency of 5S rRNA transcription from these mutant templates was determined relative to a tRNA internal control template provided by R. Devlin and M. Khosla. The results show that template sequences immediately 5' to the 5S rRNA coding region are required for efficient transcription in vitro. The C. briggsae genome contains two distinct 5S DNA repeat families ( 1.0 and 0.7 kb HindIII fragments) which are organized in mutually exclusive tandem clusters. The 1 kb 5S DNA repeat is efficiently transcribed in the C. while the 0.7 kb 5S DNA repeat appears to be transcriptionally inactive. These results suggest that C. briggsae may differentially express these two 5S DNA repeat families. The relative efficiency of 5S rRNA transcription from the wild-type C. briggsae 1 kb 5S DNA repeat and its 5' deletion derivatives in the C. confirms the requirement for 5' flanking sequences noted above. Comparison of C. riggsae sequences flanking the 5S rRNA coding sequence carried by the 1 kb 55 DNA repeats reveals the presence of a 13 bp conserved sequence. Deletions affecting the integrity of this putative regulatory element are transcribed at approximately 1/10 the wild type level. The figure below shows the location of this element and the position and transcriptional efficiency of deletion templates affecting this region.