Worm Breeder's Gazette 9(3): 40
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
While studying the gene structures of spontaneous mutations
affecting the unc-54 gene, we have identified a substantial collection
of spontaneous deletions. We have cloned and sequenced the novel
junctions of 13 of these mutations. Our goal is to determine if the
mechanism of spontaneous deletion mutagenesis in eukaryotes is similar
to that in prokaryotes. Spontaneous deletions in prokaryotes usually
have endpoints within sequences which, in the wild-type gene, are
short (5-10 bp) direct repeats; one of the repeats remains at the
novel junction. The homology that these sequence repeats provides is
important for formation of the deletions; such deletions occur at much
lower frequencies in recA strains. We find that, unlike bacteria,
sequence repeats are not a consistent feature of the unc-54 deletion
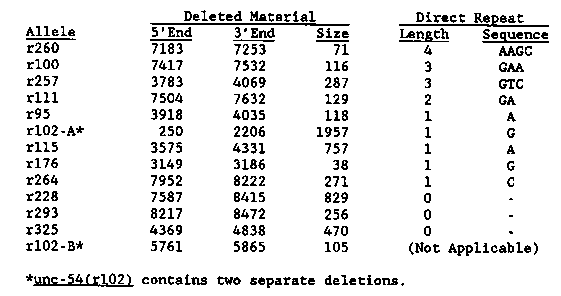
breakpoints. The table below summarizes the location and sequences of
unc-54 deletions. The base coordinates are those of Karn et al. [PNAS
80:4253(1983)].
{Figure 1}
With one exception, each of the deletions is simple, in that a
contiguous block of nucleotides is missing. No unexpected sequences
are present at the novel junctions, nor are there any rearrangements
at or near the termini. The one exception (r102-B) contains
approximately 20 bp. of unexpected DNA inserted at the novel junction.
The inserted DNA is derived from a nearby region of the unc-54 gene (
the region around base 5160). The source of the inserted material is
not deleted in r102; thus, the insert represents a displaced
duplication.
The number and sizes of direct repeats is close to that expected for
deletions having completely random endpoints. The probability of
direct repeats occurring purely by chance can be calculated from the
known base composition of unc-54. The table below compares our
observed frequency distribution to that expected for random endpoints
of twelve deletions.
{Figure 2}
The wild-type sequence of unc-54 contains a very large number of
direct repeats. For example, there are over 150 pairs of 9 bp. direct
repeats in unc-54. Deletion of the material between these repeats
would generate an unc-54 mutation that we would surely have detected
in our screens. Yet, the largest direct repeat among our deletion
endpoints is four base pairs. We conclude that direct repeats do not
contribute to spontaneous C. s in a manner
analogous to that of prokaryotes. We have been unable to identify any
consistent feature of the deletion breakpoints. We have looked for
breakpoint consensus sequences, nearby palindromes, stem-loop
structures, dinucleotide and trinucleotide frequencies, Tc1 insertion
sites, topoisomerase I and II cleavage sites, Z-DNA potential, and
sequences related to the deletion associated sequence of Dibb et al.
[J.Mol.Biol. 183:543 (1985)]. The only consistent feature of our
deletions is that they tend to be short (average size about 400 bp.).
We have sequenced one spontaneous mutation that is not a deletion.
unc-54(r274) is a G-->T transversion at nucleotide 4401, generating a
UGA nonsense codon.