Worm Breeder's Gazette 9(3): 23
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
In the last Gazette (Vol. 9, No. 2), we reported putative Tc1
mutations in a novel dumpy gene, dpy (rh1007), and in unc-44,an axonal
outgrowth gene. The analysis of these genes should yield insights into
the molecular interactions underlying morphological and neural
development.
Southern blots of the dpy (rh1007) DNA probed with Tc1 reveal only
one obvious band (at 2.7kb for Hind III digests) above those observed
both in the Bristol strain and in a spontaneous revertant. This result
indicates the initial insertion event took place either in the Bristol
chromosome or in a region of Bergerac chromosome devoid of Tc1
elements. To further demonstrate the association of the mutant
phenotype with the unique Tc1 insertion. DNA from strains produced by
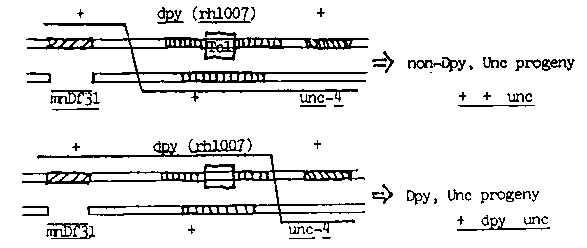
three factor crosses with flanking markers (a dpy-10 deficiency mnDf31
on the left and unc-4 (e120) on the right, see figure) has been
isolated and subjected to Southern analysis. Recombination to the left
of rh1007 should produce nondumpy, uncoordinated offspring which lack
the unique 2.7kb Hind III fragment. Conversely, progeny which are
dumpy and uncoordinated should only arise from a crossover event
between rh1007 and unc-4 and should retain the 2.7kb fragment. Six
recombinants thus far compared fit the expected pattern. The three of
these which are non-dumpy and uncoordinated strains lack the 2.7kb
fragment, whereas the three which are both dumpy and uncoordinated
retain this fragment upon Southern analysis. Therefore, the dumpy
phenotype co-segregates with the Tc1-containing 2.7kb restriction
fragment. Mutant worm DNA migrating at this molecular weight on
agarose gels is currently being isolated and cloned into the lambda
vector Charon 30.
{Figure 1}
In the case of unc-44, the six or more extra bands associated with
the mutation suggest that the initial insertion event which generated
the uncoordinated phenotype took place in the Bergerac chromosome.
Mapping the adjacent transposons is required in order to determine
which of the many Tc1 elements produces the mutant phenotype. Toward
this end, recombinant strains have been constructed from two markers
close to the unc-44 locus (bli-6 and unc-5), and DNA is being isolated
from them for Southern analysis.