Worm Breeder's Gazette 9(3): 18
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
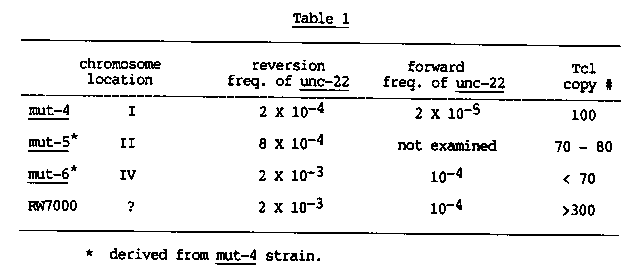
In a previous newsletter (WBG 9, #1;25) we reported experiments on a Bergerac/Bristol mixed strain, called RW7037 in which a single locus located near unc-37 I was responsible for most germ-line transposition and excision of Tc1. In further experiments with derivatives of this strain we have shown that most likely this mutator activity can itself transpose, and in the process have obtained a low Tc1 copy number strain with high mutator activity. After a further backcross of RW7037 [ genotype: mut-4(st700) I; unc- 22(st136::Tc1) IV ] with Bristol males we recovered a strain RW7080 that behaved as if it contained a second mutator, i.e., replacement of chromosome I with the Bristol homolog did not result in loss of mutator activity as it had with the parental RW7037 line. An active strain with a Bristol LGI was established and used to map this second mutator activity. In crosses with dpy-10 / + males, replacement of LGII with the Bristol homolog resulted in loss of activity: 0/26 dpy- 10 ;nc-22(st136::Tc1) lines yielded revertants in approximately 104 animals. In the reciprocal experiment, retention of the RW7080 derived LGII by elimination of dpy-10 led to retention of the mutator activity : 10/11 unc-22(st136::Tc1) Iines reverted in 104 animals. This new mutator linked to dpy-10 was designated mut-5(st701) II. A three factor cross was carried out to confirm this result : 10 + rol-1 chromosomes were recovered from dpy-10 + rol-1 / mut 136::Tc1) parents. Seventeen of the lines homozygous for the + rol-1 chromosomes retained the mutator activity; 2 did not. We conclude that during the cross to obtain RW7080 either a cryptic factor on LGII activated or the mutator activity itself transposed. These results led us to look more directly for mutator transposition, possibly into the unc-22 locus itself. We began with an RW7037 derived strain of the genotype dpy-5(e61) 00) in which the unc- 22(st136::Tc1) allele had been replaced by a Bristol LGIV. We isolated 26 spontaneous twitchers in this background and then replaced the dpy-5 me in each of the 26 established lines by the wild type Bristol homolog. In 24 cases this produced a stable strain. However, in 2 cases the unc-22 allele remained unstable after elimination of mut-4 from the background, demonstrating that the mutator activity was no longer linked to chromosome I. More detailed examination of the strain RW7096 harboring the new unc-22(st192::Tc1) allele shows it has a new mutator, designated mut-6(st702), which maps on LGIV, near or left of dpy-13 ( Of 17 dpy-13 tablished from dpy-13 / mut-6 192::Tc1 ) parents, 16 failed to revert in 10 animals; the other gave an Intermediate reversion frequency ). Table I lists some of the properties of the strains carrying these 3 mutators. In each case the principal activity can be mapped to a single region of the genome, and yet the activity varies from strain to strain, or mutator to mutator. In one case we have a strain RW7096 which has activity comparable to our original BO strain in which we had obtained evidence for a polygenic origin of mutator activity ( Genetics 108:859). If the new mutators do represent transposition of mut-4, the difference in activity of the derived and original strains is intriguing. It may be a background effect, perhaps reflecting a negative correlation between the mutator activity and the overall Tc1 copy number. Or it could be the result of changes in the mutator activity itself, perhaps reflecting a position effect on expression. In any case the strain RW7096 and its unc-22(+) derivative RW7097 are proving useful in the isolation of spontaneous mutations in a healthy low Tc1 copy number background.