Worm Breeder's Gazette 9(2): 82
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
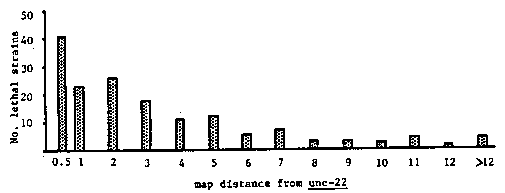
We have isolated lethal mutations in a screen to determine the distribution of essential genes on LGIV (right) in the region balanced by nT1. Previous work in the 1.5 m.u. sDf2 region has identified 20 essential genes (Rogalski, T. M. and Baillie, D. L., 1985, MGG 201:409-414). Sixteen of these lie within 1 m.u. to the left of unc- 22. unc-22(s7)unc-31(e169)/nT1(IV);+/nT1(V) hermaphrodites were exposed to .012M EMS for four hours and 3398 F1's were screened for the absence of Unc-22Unc-31 progeny. 303 lethal strains were isolated. Of the 283 strains analyzed so far, 164 lethals map to LGIV and these were retained. This data, together with data from a similar screen (Donati, L.M. et al, CSH C. 1985, p. 38), shows a frequency of lethal mutations recovered over nT1 of .078, with .044 on LGIV and .034 on LGV. The distances of LGIV lethals from unc-22 were also determined by two-factor mapping. Results show a cluster around unc 22. This parallels the distribution of visibles on the 1985 C. ese lethals are now being left- right positioned relative to unc-22 by employing a set of three deficiencies which define seven zones (sDf2, sDf21, and mDf7, which was kindly supplied by Teresa Rogalski). In the figure below, the height of the bars corresponds to the number of lethal alleles on LGIV which map in the 1 m.u. interval ending with the distance indicated. [See Figure 1]