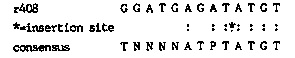Worm Breeder's Gazette 9(2): 39
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We have been analyzing spontaneous unc-15 alleles (r404, r405, r406, r407, r408) isolated from a mixed Bristol/Bergerac strain containing unc-105(n490) by Phil Anderson and Amy Bejsovec (Madison, WI). These unc-15 alleles revert spontaneously to wild type and have better muscle organization and better movement than the null e-1214. We used a lambda clone containing unc-15 specific sequence, kindly provided by Hiroaki Kagawa (Okayama U, Japan), to probe Southerns. Using several different enzymes we observed RFLDs that were 1.6kb larger in r408 than in an isogenic revertant. In order to determine if Tc1 was the cause of the mutation in r408 we cloned and sequenced this region of unc-15. The mutation was indeed due to Tc1 and the insertion site in r408 has homology to the consensus sequence derived by Ikue Mori et al (see this newsletter). [See Figure 1] The 600bp of DNA flanking the insertion site is not AT rich nor does it contain any long 0RFs with the predicted alpha-helix protein structure. unc-15 specific message is present in total and poly-A selected RNA from mixed worm populations in all of these mutants. The unc-15 message is the same size and in roughly the same amount in mutants, isogenic revertants, and N2, and there is no Tc1 hybridizing sequence in the unc-15 message. At the protein level paramyosin is easily detected on SD6 gels of whole worm extracts from these mutants. However, the amount of full length paramyosin detected by polyclonal and monoclonal antibodies in these alleles is reduced several fold from wild type levels. Just how the insertion of Tc1 into the unc-15 region produces these weak alleles is not known. From this preliminary data Tc1 appears to be reducing the amount of paramyosin produced. The DNA sequence flanking the insertion site in r408 is not intron or exon but may be in 5' or 3' regions of the gene. Further analysis of all of the alleles will be necessary to explain these observations.