Worm Breeder's Gazette 9(2): 31
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
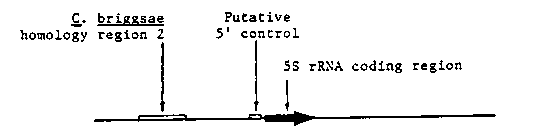
We previously reported C. elegans 5S rRNA gene structure, and demonstrated accurate, RNA pol III-specific transcription of the cloned 5S DNA in a homologous embryonic cell free extract. We are currently manipulating this system to identify template sequences and extract components required for efficient transcription. Here we report on the identification of potentially important, cis-acting template sequences. We have constructed a set of 5' and 3' deletions of a transcriptionally active 5S DNA repeat, and are examining their transcriptional activity in the nematode cell free system. Preliminary results suggest that sequences upstream of the 5S rRNA coding region are required for transcription (see figure). In contrast, X. laevis 5S rRNA transcription (in a homologous extract) is relatively insensitive to 5' flanking sequence. As an alternative way of identifying functionally important 5S DNA sequence elements, we have isolated and are characterizing members of the C. briggsae genomic repeat families encoding 5S rRNA. Preliminary analysis identifies two 120 bp regions which are highly conserved between the C..elegans and C. briggsae 1 kb 5S DNA repeats. One of these conserved regions corresponds to the 120 bp 5S rRNA; the other is located 300-180 bp upstream, in a transcriptionally dispensable region of the C. elegans repeat (see figure).