Worm Breeder's Gazette 9(2): 16
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
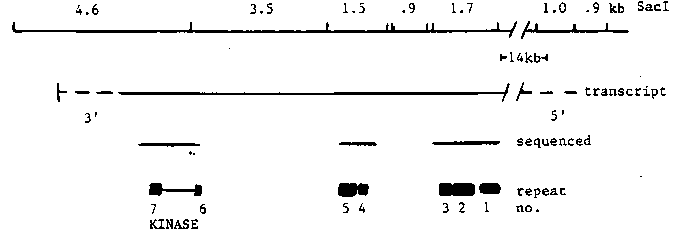
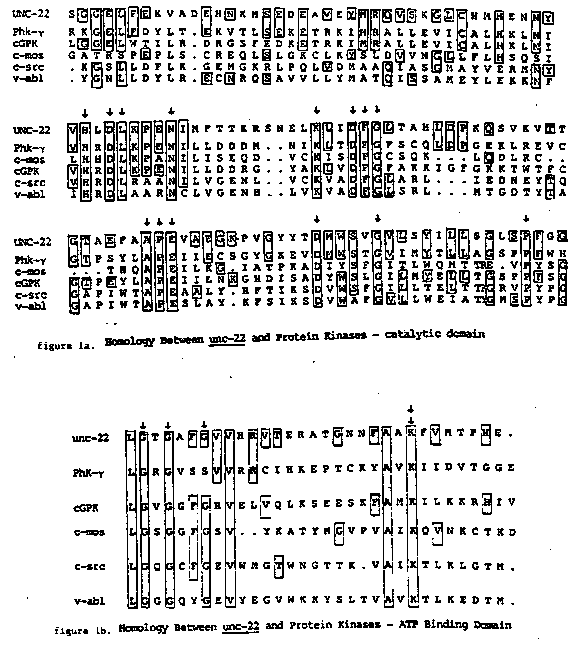
Using a probe from the 3' end of the unc-22 region, a partial cDNA for unc-22 was isolated from Barbara Meyer's lambda gt10 cDNA library. The insert is a 1.6 kb Eco RI fragment which lacks a poly A tail. By comparing this cDNA to an unc-22 genomic clone, we have determined that the cDNA lies approx. 1.5-2.0 kb upstream of the probable 3' end of the unc-22 14 kb mRNA. Both strands of this cDNA have been sequenced and an open reading frame in the direction of transcription with the biased codon usage described by Blumenthal et al. (WBG 8(1) :36) has been found. To our surprise, a search of the protein sequence data bank revealed homology to protein kinases. This portion of unc-22 is roughly 30% homologous to a stretch of about 260 amino acids comprising the catalytic domains of 3 known serine-threonine protein kinases and 10 tyrosine kinases of oncogenes (Hunter and Cooper, 1985, Ann. Rev. Biochem. 54:897). A portion of this region is shown in figure 1a. Most residues (18/24) conserved in all protein kinases are found in unc-22 and these are marked with arrows. Among these is the ala pro glu trio, likely to be important for catalysis. unc-22 also shows strong homology to the presumed ATP binding domain, which is found at the amino terminal portion of the 260 residue region (see figure 1b). This includes a lysine (marked with 2 arrows), which in the v-src protein and bovine cAMP dependent protein kinase can bind an ATP analog resulting in loss of kinase activity. In unc-22, 16 residues to the amino terminal side of this lysine is the sequence gly X gly X X gly (marked with arrows in figure lb). This glycine block is found in an analogous position in all known protein kinases and has also been observed in a number of other nucleotide binding proteins. In addition, unc-22 sequence has been analyzed from three Tc1 insertion sites and a 1.1 kb restriction fragment. A total of 4.3 kb, distributed over an 8 kb region, has been accumulated (see figure 2). We have found seven examples of a 91 residue segment. This repeat is found at the amino acid but not the DNA level. For 3 of these repeats, homology extends further, although less strongly, for a total of about 180 residues. Spacing between repeat units varies from 9 to 73 amino acids. Interestingly, 2 of these repeats bracket the putative kinase. Within the most highly conserved first 91 residues of the repeat, the same 29 residues are found in nearly the same position in at least 4 of the 7 repeats and 7 out of 29 of these residues are the same in all the repeats. Eight of the 29 conserved residues are proline. This proline richness (about 9% overall) seems to rule out alpha helical structure. No sequence homologies to known proteins were found.