Worm Breeder's Gazette 9(1): 42
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
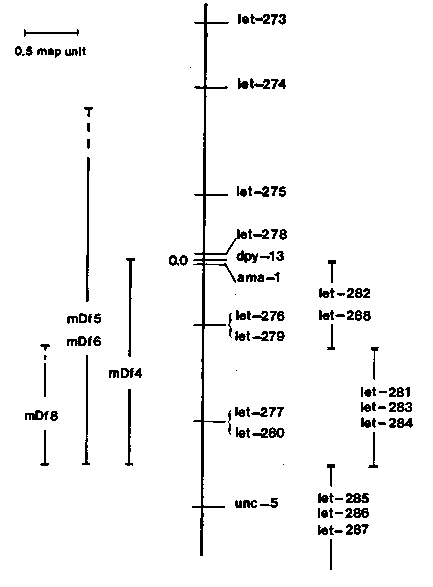
In the last issue of the WBG (Vol. 8, No. 3) we described the isolation and identification of six 'lethal' alleles of ama-1 IV, a gene encoding a subunit of RNA polymerase II (Sanford, Golomb and Riddle, J. Biol. Chem. 258:12804-12809, 1983). The six ama-1 mutations arrest development at various stages when homozygous. Two are embryonic lethals, three are adult steriles and the remaining mutation is sub-lethal (i.e. m251 hermaphrodites grow slowly and have a reduced brood size). In addition, one of the sterile mutations is temperature-sensitive. At 15 C and 20 C hermaphrodites homozygous for m238 produce some progeny (20) while at 25 C they are sterile. We are currently screening for additional mutations affecting the ama-1 gene and have isolated three new alleles, all of which are early larval lethals. The two embryonic lethal alleles of ama-1 have been further characterized by observing arrested embryos obtained from dpy-13(e184) ama-1 (m118mx)/+++ hermaphrodites using Nomarski interference- contrast microscopy. Both of these mutations appear to arrest development late in embryogenesis. Although the actual stage varied with both alleles, most of the embryos characterized were blocked during morphogenesis, between the lima bean and pretzel stages. A genetic characterization of the LGIV region around amn-1 has identified 16 essential genes in an interval of 4.5 map units (see the accompanying Genetic Map). Three of these genes are represented by more than one allele (let-272 and let-275 have two alleles; let-276 has six). In addition, this analysis has also identified three gamma- ray-induced deficiencies that include ama-1. The smallest deficiency, mDf4, is approximately 1.5 to 2.0 map units in Iength and its Ieft endpoint is located between let-278 and ama-1, an interval of 0.1 map units. Nine of the essential genes are included in mDf4. The mDf4 deficiency has been used to identify a recombinant DNA clone carrying the ama-1 gene (Bird and Riddle, this issue). The mutations that define the 16 essential genes around ama-1 exhibit a wide range of lethal phenotypes; from embryonic to maternal- effect. Eight of the genes are represented by sterile mutations, and all but one of these are located to the right of dpy-13.[See Figure 1]