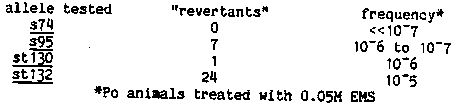Worm Breeder's Gazette 9(1): 22
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We have expanded our collection of missense alleles of unc-54 that suppress unc-22 induced twitching to a total of 17 unc-22 suppressors. These mutations can be split into two classes, an s74-like class with t4 members (see Cell 29, 773-781) and a new class with 3 members, st130, st132 and st148. This new class of mutations also act as dominant suppressors of unc-22(s12) and unc-22(st137::tc1), and a worm heterozygous for any of these mutations moves like N2 and has normal muscle structure. However, an animal homozygous for st130, st132 or st148 is slow, does not lay eggs, and the muscle cells exhibit significant A-band disorganization. It is this last trait that distinguishes this new class of mutation from mutations of the s74- like class which affect movement but have little, or no affect on A- band organization. We have carried out an extensive series of reversion experiments using s74, s95, st130 and st132 to see if there are either inter- or intragenic sites that interact with these mutations. The results are summarized below. [See Figure 1] Only st150, an s95 revertant, proved to be an extragenic suppressor and it is probably a sup-3 allele. The other revertants are the result of intragenic events. The frequency at which intragenic revertants of st132 occur as compared to s95 (a member of the s74-like class) further suggests that st132 defines a new class of mutation in unc-54. The st130 revertant and 2 of the st132 revertants are partial revertants while the other st132 revertants are full revertants to wild type. This latter set has been divided into 2 groups - of 13 tested mutations, 9 are dosage dependent, ie st132/st132, st132 m/st132 +, and st132 m/st132 m have 3 discernable movement phenotypes, and 5 are dosage independent, ie, restoration of wild type movement is fully dominant. The dosage response and the high reversion frequency of st132 indicates that compensatory second-site mutations in unc-54 can occur. The s95 intragenic mutations are full revertants and 5 of these were analyzed at the DNA level. The s95 mutation is at position 2340 in the nucleotide sequence and changes CCA (gly) to ACA (arg) (J.M.B. 183, 543-551). Fortuitously s95 disrupts a HpaII site (CCCC to CCAC). Of the 5 intragenic revertants examined by Southern analysis, only one restores the HpaII site. After cloning and sequencing the other 4 revertants we were surprised to find that all 4 have the same nucleotide change at position 2342 thus altering ACA (arg) to ACT (ser) . Our inability to find compensatory mutations in another part of unc- 54 suggests that s95 may disrupt interactions with a molecule other than myosin. The proximity of s95 to the putative ATP binding site ( EMB0 1, 945-951; P.N.A.S. 80, 4253-4257) makes ATP a likely candidate. We have sequenced 3 other mutations located in the S1 region of the myosin heavy chain. The null allele e1420 has a change at nucleotide 3960 involving CAA (gln) to UAA (ochre). Two other mutations from the s74-like class, s77 and st135, have been located. The S1 region of st135 has 2 base changes, one at nucleotide 3855 changing CAC (his) to TAC (tyr), and a second at position 3640 changing GCC (ala) to ACC ( thr). Since the (his) to (tyr) substitution is the least conservative change we suspect this is the important alteration. The change in s77 is at position 4272 and alters GGA (gly) to AGA (arg) which is particularly intriguing because the amino acid replaced is only 3 residues from thiol-1. The active thiol region has long been suspected of affecting ATP binding by the myosin heavy chain. For those up on their unc-54 fine structure map you will note that the sequence data for e1420 and s77 shows a different order than was determined by genetic mapping. In our genetic study the left/right position of these two alleles was determined on the basis of a single event, and that 'recombinant' lacked any flanking markers. Presumably then the genetic data is in error.