Worm Breeder's Gazette 8(3): 84
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
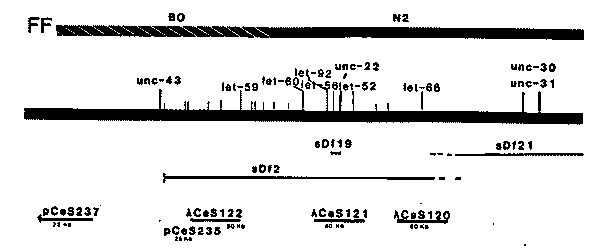
The region around unc-22, flanked by unc-43 on the left and unc-31 on the right, has been intensively studied in our laboratory over the last eight years using conventional genetic analysis. We have now embarked upon a molecular analysis of this same region. We have at present identified and recovered about 300 kb of DNA that lies within this two map unit interval. Tc1 induced restriction fragment length differences closely linked to unc-22 were identified in the N2 and BO strains. DNA was prepared from a strain constructed by serial backcrossing of N2/BO heterozygotes to N2 males (where the BO chromosome IV was marked with a EMS induced unc-22 allele). After each cross the unc-22 marked BO chromosome was recovered homozygous. This procedure was repeated six times. A lambda gt10 EcoRI library was constructed and subsequently screened for Tc1-containing sequences. These phage were isolated and their genomic fragments subcloned into pUC13 or pUC19 plasmid vectors. The flanking sequences were recovered by EcoRV digestion followed by blunt end ligation and recloning of the Tc1 deleted plasmid. These plasmids provided convenient probes for the mapping of the associated N2/BO RFLD. Three factor mapping was carried out using the marker pairs unc-43 unc-31 to map the plasmid-defined RFLD. The resulting newly-defined molecular sites are shown on the accompanying figure. The plasmids which defined each site were then used to isolate genomic clones from our lambda Charon 4 library (prepared by T. Snutch). About 25-30 kb of flanking sequence was recovered for each probe. These lambda phage were in turn checked for overlap with existing cosmids in John Sulston's genomic mapping collection. [See Figure 1]