Worm Breeder's Gazette 8(3): 82
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
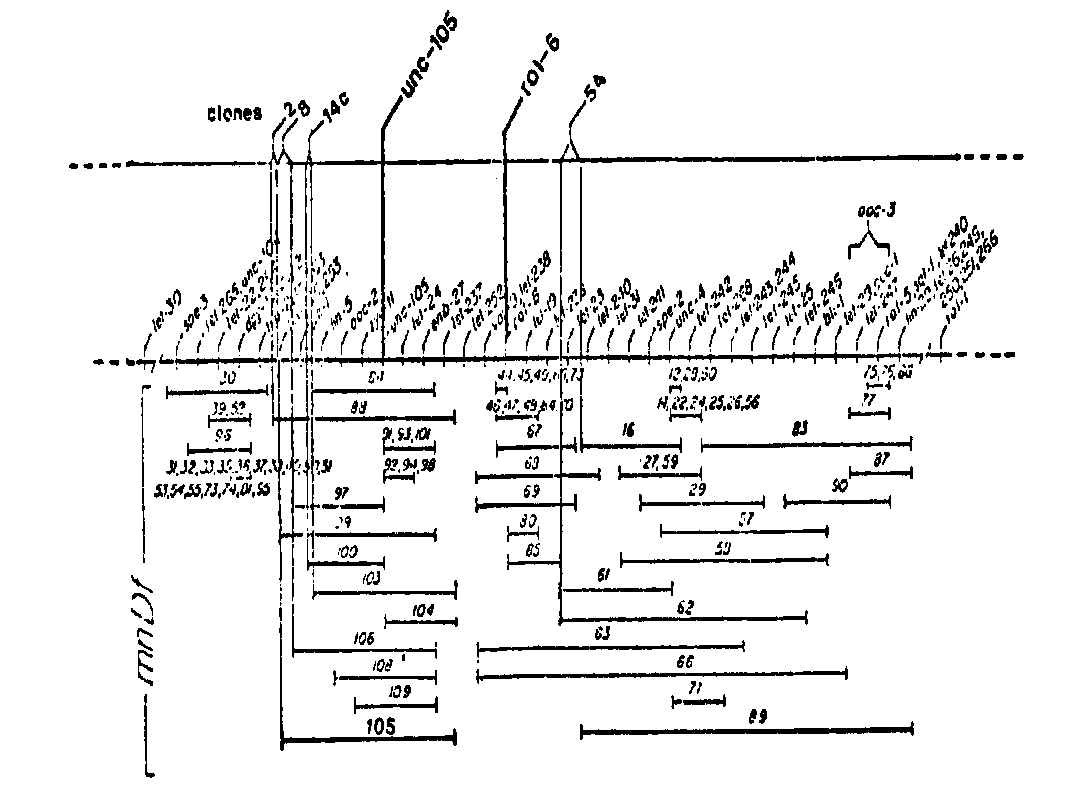
We are attempting to clone unc-105 and rol-6, both of which have been established to have wild-type null phenotypes. By repeated genetic backcrosses, we have constructed strains almost entirely derived from the C. elegans Bristol strain except for a small region of Bergerac chromosome II corresponding to the region to which both unc-105 and rol-6 have been mapped. We have localized 29 Bergerac- derived Tc1 elements within this region and have cloned in phage lambda-gt7 seven that map close to unc-105 and rol-6.We have mapped the four clones closest to unc-105 and rol-6 with respect to a series of LGII deficiencies. (Deficiency strains and the deficiency map shown below have been kindly provided to us by Chris Sigurdson and Bob Herman. The distance represented on this map corresponds to about two genetic map units, which we estimate to be approximately 600 kb of DNA. ) These clones were localized by Southern hybridization experiments using DNA isolated from Bristol strains heterozygous for different deficiencies and using the phage clones as probes. Strains of genotype mnDf unc-4/C1 unc-52 were grown on agarose plates for a number of generations, and DNA was isolated from the population of animals present, i.e., from a mixture of deficiency homozygotes (dead eggs and/or dead larvae), wild types and C1 homozygotes. Since C1 homozygotes are sterile and deficiency homozygotes are inviable, we assume that both LGII chromosomes are represented approximately equally in the DNA preparations. We have probed DNA both from N2 and from different deficiency strains with our phage clones. In each case, Southern hybridization experiments have revealed one novel band in addition to those Tc1-containing bands normally seen in Bristol strains; this novel band presumably corresponds to the unique sequence DNA surrounding the Tc1 element in the cloned probe. For cases in which the DNA of a deficiency strain resulted in this band's being of reduced intensity relative to the Tc1- containing bands (and compared to a lane containing N2 DNA), we concluded that the clone used as a probe mapped within that deficiency. The map locations of four of our clones are shown below. [See Figure 1]