Worm Breeder's Gazette 8(3): 80
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We have previously described a protocol by which we mapped Bergerac
Tc1 elements near lin-14 X (Newsletter Vol. 8 No. 1). We reported
then that three Tc1 elements mapped to within 0.2 map units of lin-14.
However, by higher resolution gel electrophoresis and Southern blot
analysis we have now found that in fact seven Bergerac Tc1 elements
map within this genetic interval. We cloned the EcoRI restriction
fragments that contain five of these Tc1 elements into phage lambda-
gt7 and used unique flanking DNA segments from each of these clones to
identify overlapping cosmids in two independent genomic C. elegans
strain N2 Bristol libraries, one constructed by Guy Benian (in Bob
Waterston's laboratory) and the other by John Sulston. Contiguous
stretches of DNA ('contigs') as defined by overlapping cosmids were
established either by traditional chromosomal walking (a total of
about 200 Kb) or by telephoning to John Sulston and Alan Coulson the
microtitre well addresses in the library of cosmids of interest. John
and Alan then kindly fingerprinted these cosmid clones (see Newsletter
Vol. 8 No. 2) and in many cases found overlaps with extant contigs
in their collection of cosmid fingerprints and contigs. By this
combination of approaches we established contigs around each of the
five Tc1 elements ranging in length from approximately 50 to 180 Kb
and totalling about 400 Kb.
Because the contigs isolated begin with restriction fragments
genetically mapping very close to each other, we expected that the
expanding contigs would overlap as new cosmids were added in the
chromosomal walk. By calculation, 0.1 map unit represents about 30 Kb
of DNA, so that the seven Tc1 elements within 0.2 map units should
have been within 60 Kb of each other. In fact, no two of the five
contigs have yet overlapped, so that the simple assumption of 300 Kb
of DNA per genetic map unit is incorrect in this genetic interval.
It was clear from these data that the resolution achieved using
recombinants within a 1 map unit interval was not sufficient to limit
or orient our chromosomal walk around lin-14. We therefore designed
an experiment in which a much rarer recombination event between a
Bristol chromosome containing two lin-14 mutations and the Bergerac
lin-14 region could be detected. This experiment allowed the Bergerac
Tc1 dimorphisms near lin-14 to be ordered relative to a recombination
point that by definition occurred within lin-14. The double mutant
lin-14 allele used was n536 n540; n536 is a semidominant allele, and
n540 is a recessive intragenic suppressor of n536. The lin-14 allele
was placed in trans to an X chromosome containing the Bristol
chromosome from near dpy-6 to the left, a region of the Bergerac
chromosome flanking lin-14(Bergerac) and containing all of the Tc1
elements near lin-14(Bergerac), and the Bristol chromosome from near
unc-9 to the right. A total of 10+E4 to 10+E5 animals of the genotype
lin-14(n536 n540)/dpy-6 lin-14(Bergerac)
eened for the n5367+ or n536/n536 n540
heterozygote phenotype of Lon, Egl, and Vul. One such animal was
found. Subsequent progeny tests showed this animal to be of the
genotype dpy-6 36)/lin-14(n536 n540), showing that
n540 maps to the left of n536 on the genetic map. DNA was prepared
from homozygous dpy-6 36) recombinant animals and
by Southern blot analysis with a Tc1 probe shown to be missing just
one of the Tc1 elements ('b,' see map below) near lin-14(Bergerac),
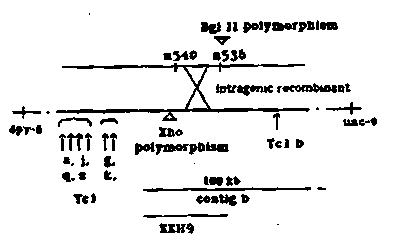
mapping this element to the right of lin-14.About 50 Kb on one side
and 130 Kb on the other side of Tc1 element b had already been cloned
by chromosome walking; therefore it was possible that even though Tc1
element b maps to the right of the lin-14 intragenic recombination
point, a cosmid at either extreme of this 180 Kb contig could map to
the left of this recombination point. Cosmids from both ends of the
contig were used as probes in Southern blot analyses of DNAs isolated
from the Bristol, the Bergerac, and the lin-14 intragenic recombinant
strains to search for Bristol/Bergerac restriction fragment length
dimorphisms that could be mapped relative to lin-14. One cosmid, KKH9,
at one end of contig b about 40 Kb from Tc1 element b, was found to
identify a Bristol/Bergerac XhoI site dimorphism that mapped to the
left of Tc1 element b, based upon the presence of the Bergerac XhoI
fragment length dimorphism in the lin-14 intragenic recombinant strain.
Therefore, the lin-14 recombination point and presumably at least
part of the lin-14 gene is located between Tc1 element b on the right
and the KKH9 defined XhoI site on the left. Cosmid KKH9 was then used
to screen DNA from 20 other lin-14 mutants for restriction fragment
length polymorphisms using a variety of restriction enzymes.
Polymorphisms were detected in DNAs isolated from mutants carrying
three lin-14 alleles, including one allele isolated after EMS
mutagenesis and two after gamma ray mutagenesis. In fact n536 (as
well as n536 n540) has an associated BglII restriction fragment length
polymorphism, which has also been detected in the dpy-6
36) intragenic recombinant strain, thus defining
the recombination point in this strain and at least part of lin-14 to
be located to the left of this BglII restriction fragment.
These data strongly suggest -- but do not prove -- that we have
cloned at least part of lin-14. We are now preparing to attempt to
definitively establish that we have cloned lin-14 by microinjecting
the clone into lin-14 mutant strains to rescue the mutant phenotype.
Unfortunately this protocol can prove but not disprove our conjecture.
[See Figure 1]