Worm Breeder's Gazette 8(1): 36
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
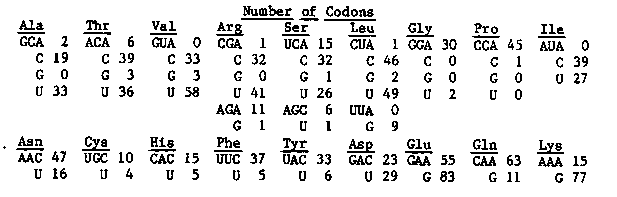
It has been found that when synonymous codons are available to code for a particular amino acid, most organisms use some codons more frequently than they use others. This asymmetry in codon choice varies considerably both in which choices are made and in the degree of asymmetry from organism to organism. Also, genes which encode abundant proteins show greater asymmetry than those for less abundant proteins [Grantham et al., NAR 9, r42 (1981)]. We present below the codon utilization table for the approximately 3.5 kb of two of the yolk protein genes which we have sequenced so far. Codon usage is extremely asymmetric in these genes. Furthermore, we have examined published sequences of the collagen, myosin, and actin genes and have found the same high degree and type of asymmetry. Although yeast and bacteria also show high (albeit different) codon asymmetries, most metazoans are not this selective about codon choice. There are several implications of these findings: 1. It is very easy to identify reading frames used for coding abundant proteins in C. elegans DNA. This can be a great aid in interpretation of ambiguous sequence data. 2. In principle, members of gene families which are pseudogenes can be so identified by comparison of codon usage between the hypothetical pseudogene and a legitimate gene. Pseudogenes will accumulate rarely used codons. This sort of analysis suggests that all five yolk protein genes are legitimate genes, even though we have been able to unequivocally assign mRNA to only two of them. 3. C. elegans seems to prefer some codons which are very rare in E. coli (for instance GGA for glycine). This may result in low level synthesis of C. elegans proteins in E. coli from clones in expression vectors. In fact, we failed to detect expression from several yolk protein DNA fragments cloned in lambda gt11 even though they were in the correct orientation and reading frame. We don't know, however, if slow translation of certain codons is responsible for this failure. The high degree of asymmetry suggests that there is strong selection for efficient translation of abundant proteins in C. elegans. In the case of yolk proteins, mutation to a less frequently used codon, almost anywhere in the gene, presumably results in reduced fecundity even though the amino acid at that position is unchanged and even though there are other genes coding for nearly identical proteins.