Worm Breeder's Gazette 7(2): 52
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
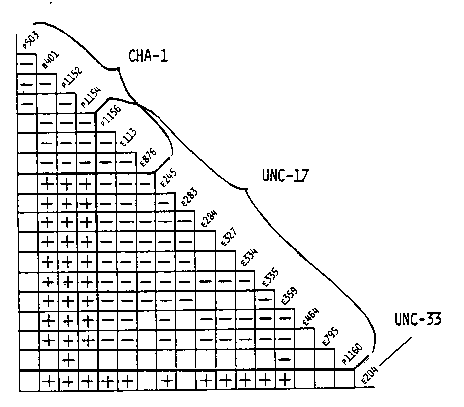
The gene cha-1 IV is the structural gene for choline acetyltransferase (ChAT) and maps very close to unc-17. Both cha-1 and unc-17 mutants have a wide range of phenotypes in common, including uncoordinated behavior (coily and jerky when backing up), slow growth, small size, and most importantly, resistance to a wide range of cholinesterase inhibitors. The evidence for two genes instead of one was: a) the cha-1 alleles that were tested all complemented unc-17 (e245) for the uncoordinated phenotype, and b) animals homozygous for e245 had normal ChAT levels. We have also been able to separate cha-1 (p1152) from unc-17 (e245) by recombination; p1152 is 0.01-0.02% to the left of e245, but such a low recombination frequency could be due to intragenic as well as intergenic events. We have now almost finished the complete complementation matrix for the uncoordinated phenotype of our cha-1 strains and the Cambridge unc-17 strains. Most of the alleles fall into one of two disjoint complementation groups, but there are three anomalies: p1156 ( originally isolated as being cha-1) and e113 and e876 (both isolated as unc-17) appear to be members of both complementation groups. A clear prediction from the complementation pattern was that animals homozygous for e113 or e876 would lack ChAT (we already knew that p1156 had very low ChAT), but enzyme assays showed that all the unc-17 strains, including e113 and e876 homozygotes, had between 60 and 150% of the wild-type ChAT activity. At the moment, then, we have two overlapping complementation groups, with a lack of uniformity in the overlap set. We have been considering various models, including a protein with two discrete, independently functioning domains, or a deletion hotspot located between two adjacent genes, or even overlapping genes. Next on the agenda is careful biochemical analysis of ChAT from the unc-17 strains. [See Figure 1]