Worm Breeder's Gazette 7(1): 80
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
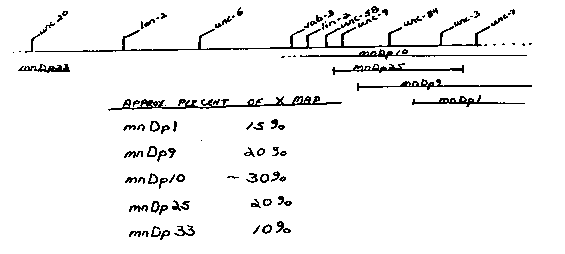
Animals homozygous for dpy-21 V are Dpy if they have two X chromosomes (X:A = 1.0), but nonDpy if they have only one X (X:A = 0.5) , irrespective of sexual phenotype. Somehow the autosomal dpy-21 locus appears to respond to the ratio of X chromosomes to sets of autosomes (X:A ratio; Hodgkin, Genetics 96:649-664, 980). I have used the X-linked duplications from Minnesota in attempts to determine whether a specific region, or simply the total amount of X chromosomal material is being counted. The basic idea is to construct mnDP(unc+)/+; him-5 unc X stocks and to look at the nonUnc male progeny which will have one X plus the duplication. If the duplication is not counted as an X chromosome (either because it is too small or because it lacks the crucial regions, these males will be nonDpy; this result is found for mnDp1, mnDp33.duplication is counted as an X chromosomes, these males will be Dpy. Both mnDp10 and mnDp25 may fall into this class, because each gives a few Dpy males, and some semiDpy males. As seen in the map and the table, both mnDp10 and mnDp25 include a region to the left of unc-9; however, they are also apparently the largest duplications, based on the map distance between duplicated markers, a crude estimate at best. Thus, dpy-21 responds to these two duplications at least partly as X chromosomes, either because they carry a region to the left of unc-9 or because they represent a substantial amount of X material. An experiment designed to discriminate between these possibilities has given only preliminary results so far. I have attempted to construct a stock that has both mnDp33 and mnDp9; neither of these by itself is counted as an X, but together they represent about as much X material as mnDp10. The last step in the construction was to mate mnDp33/+; him-5 unc-20/0 males to mnDp9/+; him-5 unc-20, odites. The males from this cross will include some nonUnc-20 nonUnc-3 males that have both duplications; all other males will be Unc-3 or Unc-20 or Unc- 20 losely resembles Unc-3). There were 6 nonUnc males and all were Dpy, suggesting that mnDp33 and mnDp9 together are counted as an X chromosome. Unfortunately, these males did not mate, so I could not make an hermaphrodite stock with both duplications. A straightforward interpretation of these results would be that in responding to the X:A ratio, dpy-21 does not distinguish between one X plus 25%-30% of an X (X:A=0.65% and 2X (X:A=l). However, this view appears contrary to Jonathan Hodgkin's observation that 4A:4X tetraploid dpy-21 hermaphrodites (X:A=l) are Dpy, while 4A:3X dpy-21 hermaphrodites (X:A=0.75) are nonDpy (ibid.). The two sets of observations can be reconciled by somewhat fancier models, for example, by assuming that the crucial X material is not homogeneously distributed along the X. [See Figure 1]