Worm Breeder's Gazette 6(1): 26
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
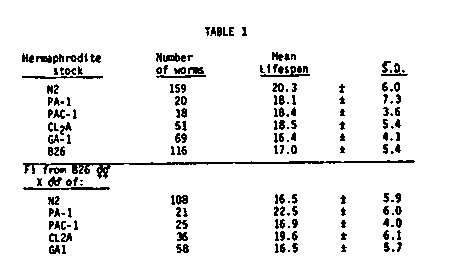
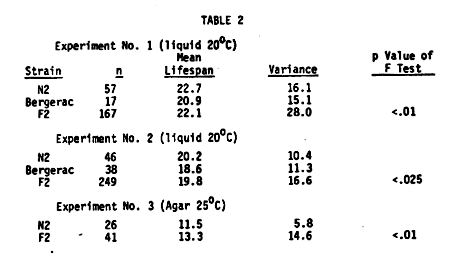
Growth Conditions: We are using lifespan as a quantitative variable for performing several sorts of genetic analyses. We have picked standard environmental conditions which optimize the brood size of hermaphrodites. These conditions are 20 C in 4 mls of S basal plus cholesterol with 10+E9 E. coli B/5 as food source and 50 worms per plate in 6 cm falcon plastic petri plates. In this range, lifespan is negatively correlated with both temperature and with E. coli concentration. These conditions are therefore far from optimal for length of life. We have also used standard NGM and OP50 at 25 C as an alternative environment for growing aging populations. Under these conditions only 10 worms are cultured on each plate. The lifespan of all populations is significantly shortened under these latter conditions. Data Storage and Analysis: We are developing a system for selection of strains of worms which display significantly lengthened lifespans when compared with normal N2 strains. Much of the analysis of lifespan is based on comparisons of survival curves for different populations. We have developed a computer based data storage and analysis system. The system stores data on a daily basis and then performs several transformations to yield data sets suitable for analysis either by SPSS (statistical package for social sciences) or by the log rank statistic. Both the SPSS which uses a modified Kruskal-Wallis test, and the log rank are non-parametric statistics developed precisely for performing survival analysis on populations of individuals. Preliminary evidence suggests that the log rank is a somewhat more powerful statistic. We are performing comparisons on these statistics now. Reproducibility: Reproducibility is tested by asking whether genetically identical sample populations generated simultaneously but independently transferred by different individuals show identical survival curves. They do. However, variability is introduced by minor changes in procedure. Sample populations of N2 established in different experiments show significant differences in lifespan. This problem is overcome by including a full battery of controls in each experiment. Inbreeding and Heterosis: Most animal species have characteristic lifespans under defined experimental conditions. C. elegans N2 also shows different lifespans for the male and the hermaphrodite. Typical mean N2 male and hermaphrodite lifespans are about 17 and 20 days, respectively. Other laboratory species previously used in analyses similar to ours show overdominance for length of life: that is, the F1 hybrid progeny of two different lab strains live significantly longer than either parental line. C. elegans does not show this effect ( Table 1). The F1 progeny of two different lab strains have lifespans intermediate to the mean lifespan of the two parental wildtype strains. Thus lifespan appears to be controlled by simple additive genes whose effects can merely be summed to predict the probable lifespan of the F1. This major difference in the behavior of C. elegans and other animal species may be due to the fact that the worm is a self- fertilizing hermaphrodite and is normally completely inbred. In sexual species significant inbreeding occurs in the laboratory environment, so that lifespans of lab populations often are shorter than lifespans of wild populations. Cross breeding of such lab populations results in a significant overdominance effect on lifespan. The lack of overdominance effects in C. elegans allows us to use length of life as a simple quantitative genetic parameter. Quantitative Genetics of Lifespan: We have established populations of worms derived from F1 individuals of a cross between Bristol (N2) males and Bergerac hermaphrodites. The F1 individuals were raised at 25 C to identify Bergerac self-fertilization progeny which are sterile at 25 C. Fertile F1 outcross progeny produced F2 progeny which were maintained at 20 C to allow normal development of those individuals carrying the temperature sensitive Bergerac gene. The F2 populations derived from such experiments consistently show an increase in variance of lifespan (Table 2). F tests on the data show that this increase is significant. We can demonstrate even greater differences by performing the survival curves on agar at 25 C instead of in suspension at 20 C (Table 2). Effects of EMS Treatment: No one has developed a strain, in any organism, with significantly increased longevity. We have examined the effects of EMS mutagenesis on length of life of the parental mutagenized animals and their F1 or F2 progeny. We find significant deviations from the unmutagenized controls in the F2 of EMS treated parents (Table 3). However, EMS treated parental worms or their F1 progeny show no significant differences in lifespan. This can be explained by assuming that the differences seen among the F2 are due to homozygosis of many sublethal genes. Screening of Potential Genetic Markers: We have measured lifespans of several ts developmental mutants, X-linked morphological and behavioral mutants as well as sexual transformers of several types. One major purpose behind these screens is to identify markers that do not affect lifespan. A partial summary of these findings is presented in Table 4. We have asked if the lifespan depression observed in one of the strains, B245, might be due to other unlinked genes which are still segregating in the B245 genetic background. Among several isolates from a backcross of B245 with N2 males was one isolate which did display normal N2 lifespans. We are currently repeating this experiment. These findings suggest that one reason for not detecting mutants which extend lifespan might be that the background of EMS induced sublethals blocks the observation of those mutants which do show an extension of life. Selection for Increased Lifespan: We are currently beginning the second round of a long term selection designed to select longer-lived strains from a Bristol-Bergerac hybrid population. A set of F2 progeny are generated as outlined above and maintained as single worms in microtitre plates. F3 progeny of individuals are saved until the longest lived worms are identified. Males are generated from these F3 plates and all the long-lived populations are then intercrossed to produce a new hybrid population. [See Figure 1] [See Figure 2] [See Figure 3]