Worm Breeder's Gazette 17(1): 60 (October 1, 2001)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Dept. of Biology, Univ. of San Diego, San Diego, CA 92110
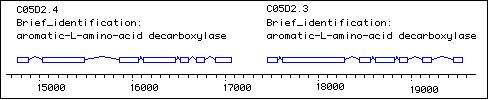
We are characterizing genes used by the serotonergic neurons in C. elegans to learn how they are regulated; among these is the bas-1 gene, which encodes a serotonin- and dopamine-synthetic aromatic amino acid decarboxylase (AAADC). Mutants in bas-1 are serotonin- and dopamine-deficient. Wildtype serotonin immunoreactivity (IR) can be restored by application of exogenous serotonin, but not its immediate precursor 5-HTP; this phenotype is consistent with the loss of serotonin-synthetic aromatic amino acid decarboxylase activity. Wildtype serotonin-IR is rescued in bas-1 mutants by injection of the cosmid C05D2 and various subclones containing two adjacent AAADC genes called C05D2.4 and C05D2.3 (see figure below). We now show that the first predicted gene, C05D2.4, is required for rescue of bas-1 and encodes the serotonin-synthetic AAADC activity in C. elegans.
We have identified the mutations in four bas-1 mutant alleles (ad446, pa4, n2948, n3008); a fifth allele (tm351) was recently generated by the C. elegans Gene Knockout Consortium with a large deletion of C05D2.4 coding sequence (thanks!). Both the tm351 homozygote and tm351/ad446 are serotonin-(-IR)-deficient. The alleles pa4, n2948 and n3008 contain point mutations in C05D2.4 coding sequence resulting in premature stop codons. The original bas-1 allele, ad446, contains a 4268 bp deletion from the second exon of C05D2.4 to the final intron of C05D2.3, so is a knockout of both predicted genes. [Vexing question: why do ad446 worms have residual serotonin-IR?] Our results are consistent with the hypothesis that mutations in C05D2.4 cause the Bas phenotype. We have also determined that a construct mutated in C05D2.4 (frameshift creating a premature stop codon) does not restore serotonin-IR to bas-1 mutants (2 independent lines) whereas constructs mutated in C05D2.3 do rescue bas-1 (frameshift creating a premature stop codon - 1 line; GFP insert into coding region - 2 lines). The smallest subclone we have tested to date that rescues bas-1 mutants has 8.8 kb containing both genes intact, with 2.6 kb upstream of the C05D2.4 start.
C05D2.4/bas-1 cDNAs are trans-spliced to SL1. We have isolated bas-1 cDNAs with RT-PCR; we have received other cDNAs from the EST project (thanks to Y. Kohara et al.) and the ORFeome project (thanks to J. Reboule et al.). Two YK clones came from the 'full-length' capped library, and each has an SL1 leader and poly-A tail, but both contained internal deletions (overall abnormality of these clones reported at ~5% - J. Thierry-Mieg et al., 2001 IWM Abstract 231). We have cloned two splice variants different from the Genefinder-predicted cDNA (the predominant form); to date we have found one of these also among ORFeome project-derived clones (also predominantly the predicted form). In one variant, a 27 bp microexon is spliced in between predicted exons 2 and 3. A second variant uses an alternative splice acceptor 60 bp upstream of the usual splice site for exon 3; this alternate splice introduces a premature stop codon in the coding sequence. It remains unclear what functional significance these splice variants may have.
The predicted genes C05D2.4 and C05D2.3 are very close together -- only 369 bp apart from predicted stop of C05D2.4 to predicted start of C05D2.3 (figure). This could indicate the two genes are transcribed together as an operon. To date, however, there is no evidence that the genes are transcribed together. Although there is an EST project partial cDNA, and we have sequenced a partial cDNA for C05D2.3, we have not yet isolated an SL1 or SL2-spliced cDNA using RT-PCR. Furthermore, C05D2.4 and C05D2.3 are not among genes identified as likely operons by microarray expression analysis (T. Blumenthal, personal communication). The function of C05D2.3 remains a mystery. Intriguingly, one of C05D2.3's 15 "nearest neighbors" in the gene expression map using the "Worm Proximity Analyzer" (http://workhorse.stanford.edu/cgi-bin/murray/nneighbors.cgi) is F12A10.3 - one of the other AAADC genes in C. elegans.