Worm Breeder's Gazette 16(5): 18 (February 1, 2001)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
| 1 | Anesthesiology Research Division, Laboratories of Cellular and Molecular Physiology, Departments of Anesthesiology and Pharmacology, Vanderbilt University Medical Center |
| 2 | Department of Cell Biology, Vanderbilt University Medical Center |
Random deletion mutagenesis provides a powerful mechanism for knocking out gene function in C. elegans. A "library" of mutagenized worms is stored frozen at -80ºC and lysates of individual worm cultures are screened for deletion mutations in specific genes by PCR. Once a mutation is detected, cultures are thawed and worms harboring the mutation are identified by further PCR screening (1).
Because mutant alleles are present at very low frequency, high survivorship of frozen worms increases the likelihood of detection and isolation of mutant worm strains. It is widely known among C. elegans workers that the rate of cooling and freezing of worms is key to good survival (2). Here, we report improved survivorship of frozen C. elegans by modification of a standard freezing protocol. Worms are frozen in an insulated cooler designed originally by Ranganathan and Reddien (http://info.med.yale.edu/mbb/koelle/protocol_list_page.html). Typically, worms are frozen by direct placement of the cooler in a -80ºC freezer. However, as shown below, worm survival is increased by pre-cooling the worms at -30ºC.
Synchronized N2 Bristol L1 larvae were suspended in M9 buffer and diluted to a density of approximately 160 larvae/10 ml. Sixty microliters of this worm suspension were then mixed with an equal amount of standard freezing medium in a PCR tube. Ten of these PCR tubes were placed among a stack of six empty freezing plates inside a fully-assembled cooler that was then placed into a -30ºC freezer. Ten similar PCR tubes were placed in the -30ºC freezer on a rack without any insulation. Finally, a second cooler with ten more PCR tubes of larvae and freezing medium was placed at -80ºC. Five hours later, both the cooler and the non-insulated tubes at -30ºC were moved to the -80ºC freezer. The next day, all thirty tubes were thawed and the contents of each tube were plated onto a separate 60 mm NGM plate seeded with E. coli OP50. Two days later, the number of live worms on each plate was determined and the data were analyzed by one-way Model I ANOVA after square-root transformation.
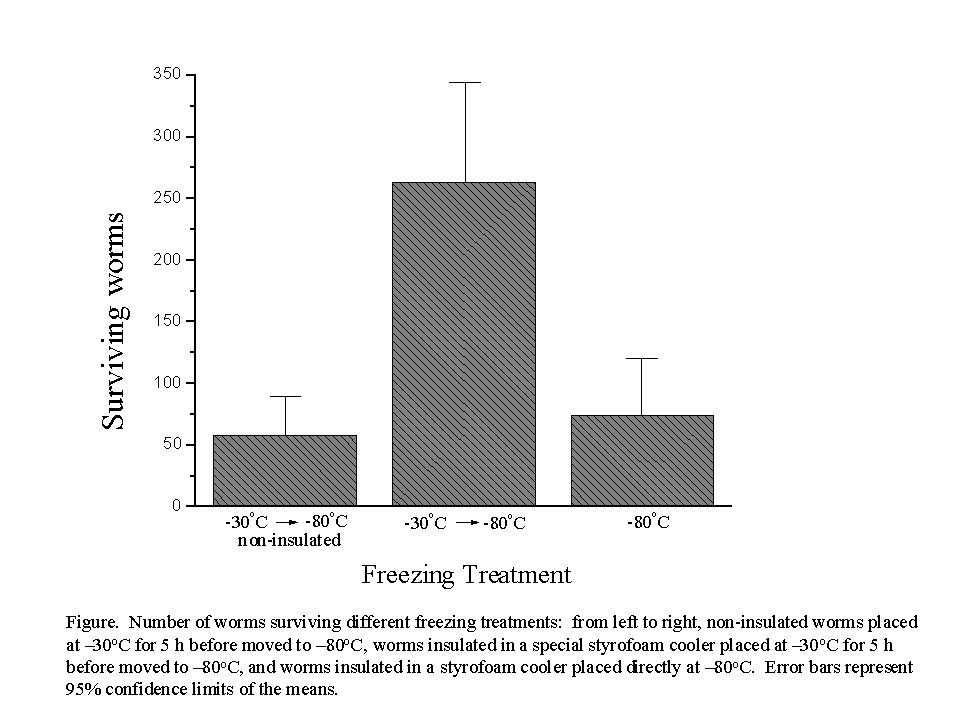
Worms frozen in the special cooler at -30ºC for 5 h showed significantly higher survival than both the non-insulated -30ºC treatment and the worms in the cooler placed directly at -80ºC (F = 16.99; df = 2, 29; P < 0.0001; Figure). The combination of the insulating cooler and the 5 h treatment at -30ºC prior to moving to -80ºC increased survivorship 3.5-fold. We postulate that the improved survivorship is due to a reduced rate of cooling in the cooler maintained at -30ºC. Workers are urged to optimize empirically their own freezing protocols to account for variation among freezers and conditions in different laboratories.
References:
1. Liu, LX, JM Spoerke, EL Mulligan, J Chen, B Reardon, B Westlund, L Sun, K Abel, B Armstrong, G Hardiman, J King, L McCague, M Basson, R Clover, and CD Johnson. 1999. High-throughput isolation of Caenorhabditis elegans deletion mutants. Genome Research (9): 859-867.
2. Lewis, JA and JT Fleming. 1995. Basic culture methods. In Methods in Cell Biology Vol. 48: Caenorhabditis elegans: Modern Biological Analysis of an Organism. HF Epstein and DC Shakes, eds. Academic Press, San Diego.
