Worm Breeder's Gazette 16(2): 30
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
| 1 | Institute of Cytology and Genetics, prospect Lavrentjeva 10, Novosibirsk 630090, Russia |
| 2 | The Netherlands Cancer Institute, Division of Molecular Biology, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands |
We are interested to use C. elegans for studies of retrotransposition processes and particularly regulation of non-LTR elements. The rte-1 retrotransposon is the most abundant element in C. elegans genome and has all structural features of an active non-LTR element. We have studied polymorphism of rte-1 in wild-type strains to estimate levels of its activity.
Southern blot analysis of 20 natural C. elegans isolates revealed some differences in rte-1 banding patterns between strains. All strains contain approximately the same number of rte-1 copies. Observed differences in banding patterns are due to insertional polymorphism of rte-1 but could also be explained by other RFLPs unrelated to rte-1.
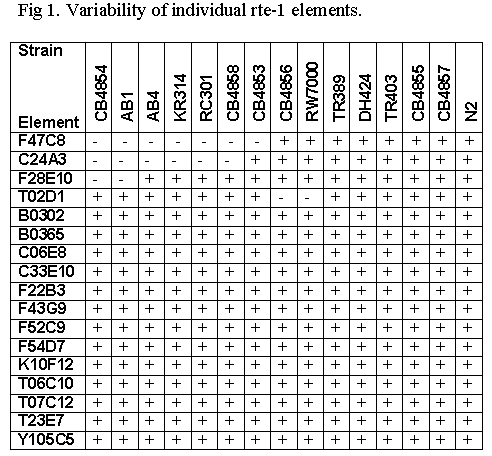
Next we have checked the presence of individual rte-1 elements in different strains by PCR with elements-specific and site-specific primers. Among 17 loci analysed for 15 strains 4 loci appeared to be variable for insertion of an rte-1 (Fig. 1). "Empty" sites have been PCR amplified and sequenced. Analysis of variable sites allowed to determine precisely changes in target sequences which take place upon insertion of rte-1 elements. Briefly, insertion of rte-1 element may cause a) deletion of several nucleotides in target sequence; b) formation of target site duplications of variable length; c) addition of tandem repeats derived from upstream target site region to the 5' end of the element; d) addition of non-templated nucleotides to the 5' end of the element. All of the above kinds of structural changes have been previously reported for other non-LTR retrotransposons and may be explained by a template switch model of retrotransposition.
Insertional polymorphism of rte-1 element observed in wild-type strains indicates that rte-1 at present is an active, capable of retrotransposition element. However the rate of retrotransposition seems to be quite low. Currently we make attempts to mobilize rte-1 in laboratory strains to detectable levels of transposition.
