Worm Breeder's Gazette 16(2): 24
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Division of Molecular Biology, The Netherlands Cancer Institute, Center for Biomedical Genetics, Plesmanlaan 121, 1066 CX Amsterdam, The Netherlands
Previously, (WBG, 16(1)), we described a confirmed set of SNPs from CB4856 that can be detected quickly by restriction digest of single worm lysate (snip-SNPs). At that time, about 100 markers had been confirmed. Currently, the snip-SNP map has almost 200 confirmed markers. Additional markers have been added in a directed fashion, with the intention of reducing the larger intervals that existed on the map. We estimate that useful snip-SNPs are present in the CB4856 at a density of more than one per cosmid.
We have developed a protocol using these snip-SNPs, based on bulked segregant analysis (Michelmore, et al. (1991) Proc.Natl. Acad.Sci. USA 88, 9828-9832), that facilitates rapid mapping of genetic loci. Single worm lysates are derived from F2 mutant and wild type cross progeny. A fraction of each lysate from all mutant F2 worms is pooled into a single bulked lysate; a wild type lysate is made from the wild type worms. These lysates then act as a templates for snip-SNP genotyping. If a marker is linked, the CB4856 form of that marker is excluded from the bulked lysate obtained from mutant worms, and is enriched in the bulked lysate obtained from wild type worms. The relative amount of one form of the biallelic marker to the other in a given lysate is a measure of how linked the marker is to the target mutation. By taking a ratio of these amounts derived from the mutant and the wild type lysates, it is possible to calculate a Map Ratio. The Map Ratio of unlinked markers is about one (the relative amount of CB4856 DNA at that point in the mutant bulked lysate is roughly the same as the relative amount of CB4856 DNA in the wild type lysate); the Map Ratio of linked markers is significantly less than one. Eighteen SNPs are checked to establish linkage to a chromosome arm, and a higher resolution set from the linked chromosome is then checked to localise the mutation to a small interval.
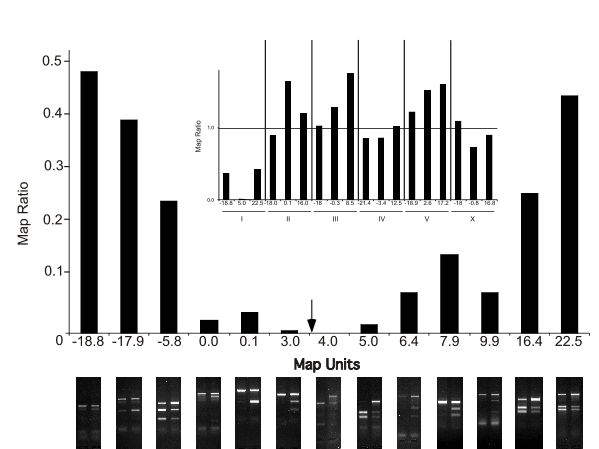
Using this approach, we have been able to map mutations to map unit resolution using a single cross of CB4856 with the mutant dyf-5. Mapping was completed within 8 hours of isolating F2’s; no visible markers were required either in cis or in trans to the mutation, thus no mapping strains needed to be constructed. In Figure 1, the inset establishes linkage to the centre of chromosome I using 3 snip-SNP markers per chromosome, and the large figure represents 13 markers dispersed over chromosome I. The arrow indicates the published map position of dyf-5. Beneath the axis are images of the BSA markers used to calculate the Map Ratio at each indicated position on chromosome I. For further fine mapping, using remaining DNA from the same animals, it is possible to select recombinants within the identified small interval. In principle, only a single cross is required to map a mutation to an interval of one or a few cosmids in a very short period of time.

While this protocol is generally useful, it has particular utility for mapping otherwise problematic mutations such as lethals, dominant mutations, polygenic traits and behavioural mutants with phenotypes that will not tolerate visible markers.