Worm Breeder's Gazette 13(4): 94 (October 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Institute of Molecular Biology and Biochemistry, Simon Fraser University, Burnaby, BC
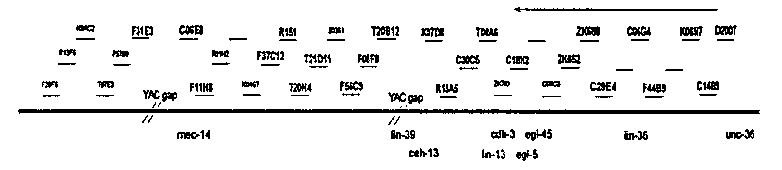
As reported in the June 1994 issue of the WBG, the sequencing consortium is rapidly completing the sequence of chromosome III. Concurrently, our lab has been generating a large number of new mutations in the region covered by the free duplication sDp3 (approximately 75% of chromosome III, left) (1). These mutations include EMS induced point mutations in essential genes and small, UV/Gamma induced deletions (see H. Stewart et al, this issue). In collaboration with the C. elegans genome sequencing consortium, specifically the St. Louis lab, our lab has begun the task of correlating the genetic and physical maps on the left arm of chromosome III. As a first step toward this goal, I have been micro injecting cosmid DNA received from St. Louis (my thanks to P. Latreille) into N2 hermaphrodites to create stable transgenic strains carrying specific sequenced cosmids. It is our intention to use these transgenic strains to cross these cosmids into various mutant strains and observe if the DNA is able to rescue the mutant phenotype. In this manner we will be able to place a large number of genetic markers on the physical map. Germline transformation is performed as per Mello et al. (1991), with cosmid DNA normally being injected at an approximate concentration of 20 ng/265l. Prior to injection, cosmid DNA is mixed with the plasmid pCes 1943 to form an injection mixture having a total DNA concentration of approximately 100 ng/265l. pCes 1943 is derived from the plasmid pRF4 (2). It contains the dominant rol-6 ( su1006 )allele (3), and has been modified to contain a kanR cassette. Transformant strains which stabily give rol-6 progeny are PCR tested to confirm the presence of cosmid DNA. It has been my observation that approximately one stable rol-6 transformant in four does not contain cosmid DNA. So far I have produced transgenic strains from 22 cosmids: C06E8 , C06G4 , C14B9 C18H2 , C29E4 , C30C5 , D2007 , F08F8 , F11H8 , F31E3 , F37C12 , F44B9 , K06H7 , K07D8 , R151 , R13A5 , T04A6 , T20B12 , T20H4 , T21D11 , ZK652 ,and ZK688 (see figure). Although there are a few cosmids left of mec-14 which I have not been able to successfully inject (due to problems with DNA availability or possible dosage effects), these transgenic strains represent a reasonably continuous stretch of DNA approximately 654 000 bp long. Preliminary crosses between lethal bearing strains and cosmid-containing transformants have yet to reveal a cosmid rescue. However, as our newly generated mutations are further mapped with respect to previously generated physical markers, the task of choosing cosmids which are likely to rescue particular mutants will become easier. [see figure] In the meantime we are making these transformant strains available to the C. elegans community. If you would like us to send you any of these strains please contact me at: dcollins@darwin.mbb.sfu.ca This research is being funded by a grant from the Canadian Genome and Associated Technologies (CGAT) (1) Collins, D., H. Stewart and D. L. Baillie (1994). WBG Vol. 13 No. 2: 66