Worm Breeder's Gazette 13(4): 67 (October 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
| 1 | MGH Cancer Center, Boston, MA. |
| 2 | MGH Cancer Center and Dept. of Pediatrics, Boston, MA. |
| 3 | MGH Cancer Center, Boston, MA. |
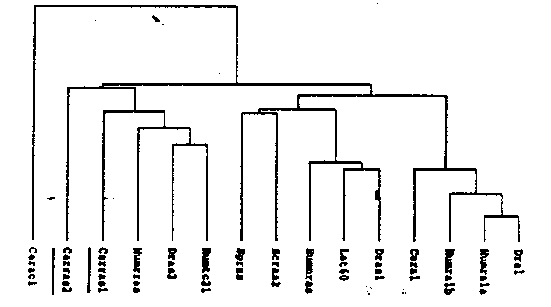
Human r-ras 1 and r-ras 2 ( TC2 1))belong to the closer relatives (>50% amino acid identity) of ras in the ras superfamily of GDP/GTP-binding proteins. They are the first members to exhibit transforming potential when mutated at some residues which render ras oncogenic and make it insensitive to GAP action (Graham & Der, 1994). These recent findings have led to current investigations of their role in human cancer. Furthermore, r-ras 1 -- by immunoprecipitation and in the yeast-2-hybrid-system -- was shown to interact with bcl-2 ,the human homolog to ced-9 (Fernandez-Sarabia & Bischoff, 1993) and has thus been implicated as a possible effector of apoptosis. There is evidence that the r-ras proteins participate in some but not all aspects of the ras signal transduction pathway involving upstream tyrosine kinases and downstream serine/threonine kinases. It has not yet been elucidated in the mammalian system (1) what alternative pathway the r-ras proteins may be utilizing and (2) what functional relevance is represented by the in vitro interaction of r-ras 1 and bcl-2 . We are trying to address these questions in C. elegans and have cloned the homologs of r-ras 1 and r-ras 2 using a degenerate PCR approach. We have screened c-DNA and genomic libraries and obtained and sequenced full length c-DNA and genomic clones of r-ras 1 and a full length c-DNA clone of r-ras 2. The genomic sequence of r-ras 2 was recently made available by the genome sequencing project. The amino acid comparison shows high homology/identity to the human proteins for r-ras 1 and r-ras 2 ( TC2 1);the location of the C. elegans proteins in a phylogenetic tree containing the closer members of the ras superfamily (including some members of Drosophila and saccharomyces) is shown below. (see figure) R-ras 1 was localized to chromosome II near lin-29 ,and r-ras 2 maps close to emb-5 on chromosome III. To obtain r-ras germline deletions, we have screened a TC1 insertion library which we constructed using the mutator strain MT 3126 (protocols kindly provided by Joel Rothman, Susan Mango and Ed Maryon), and have isolated transposon insertions in r-ras 1. We arc currently in the process of sib selection to purify the strains. To get some first appreciation of a functional role of r-ras towards apoptosis versus growth stimulating properties, we have also started to inject a r-ras 1 heat shock promotor expression construct to generate strains in which r-ras can be overexpressed This additional approach has been chosen since redundancy may be expected in the ras related protein families and thus the knockout of one of the proteins may not give clear results. We will screen the overexpressing strains for (1) apoptosis and (2) muv phenotype. In collaboration with Bob Horvitz's laboratory r-ras GST fusion proteins will be generated to test the in vitro interaction with ced-9 . Finally, we are constructing r-ras 1 and r-ras 2 promotor expression vectors with GFP/betaGAL to define the expression patterns of both genes.