Worm Breeder's Gazette 13(4): 56 (October 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Department of Biology, Indiana University, Bloomington, Indiana, 47405
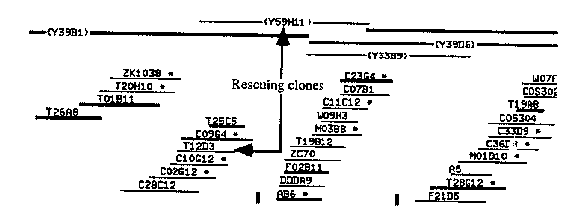
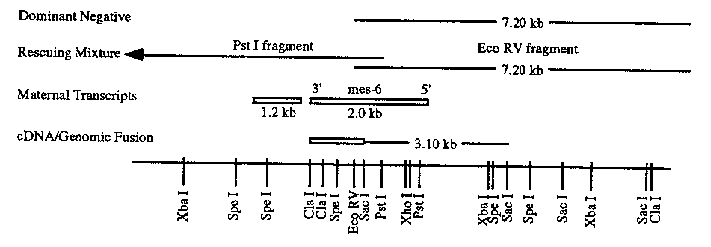
mes-6 was identified through screens for maternal effect sterile loci. Its phenotype is similar to that of mes-2 , mes-3 ,and mes-4 .Homozygous mutants at these loci produce progeny with few germ nuclei and no gametes. We are cloning these genes to better understand their function. mes-6 is located between unc-24 and fem-3 on LG IV. The physical map of this region is shown in Figure 1. Using transformation rescue, we were able to identify a YAC ( Y59H11 )and a cosmid ( T12D3 )that contain mes-6 .Single restriction fragments of T12D3 did not rescue but a curious phenomenon was observed: a 7.2 kb Eco RV fragment induced an array-dependent dominant sterile phenotype. We hypothesized that this may be due to a portion of the gene acting in a dominant negative manner. A mixture of the Eco RV fragment and an overlapping Pst I fragment rescued mes-6 .These fragments were used to probe a Northern blot. Only one transcript hybridized to both restriction fragments. This 2.0 kb transcript is enriched in strains containing oocytes ( N2 and fem-2 lf) relative to those without a germ line or containing only sperm ( glp-4 and fem-3 gf). This maternal expression is consistent with the mutant's maternal-effect phenotype. The Pst I restriction fragment detected another 1.2 kb maternal transcript. The cDNA corresponding to this transcript maps very close to the 2.0 kb transcript and they may define an operon. In order to prove that the 2.0 kb transcript is mes-6 ,we translationally fused a 3.1 kb Sac I genomic fragment, containing the 5' half of the gene and upstream sequences, to a 1.0 kb cDNA fragment containing the 3' half. This construct rescues mes-6 .The data are summarized in Figure 2. Sequencing of both mes-6 and the neighboring gene is in progress. (see figures)