Worm Breeder's Gazette 13(4): 18 (October 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Gene Library Lab, National Institute of Genetics, Mishima 411, Japan e-mail: ykohara@ddbj.nig.ac.jp
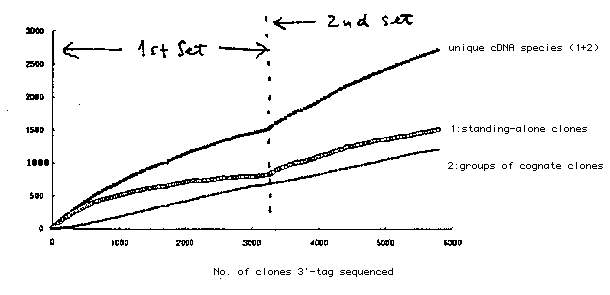
(1) First set of cDNA clones: A set of some 4,400 cDNA clones selected from a library of size-fractionated (=>2kb ) cDNA made from a mixed-stage population of him-8 strain have been analyzed with respect to (a) tag-sequencing from both ends, (b) mapping onto the genome, and (c) surveying expression pattern as described (WBG 13-#2, p.20). All the information on (a) and (b) has been sent to ACEDB and the tag sequences are already in the public DNA database. (a) Tag sequencing: Out of the clones, 3,194 clones gave clean 3'-tag sequences, which were classified into 1,508 unique species (831 standing-alone clones and 677 groups of 2-23 clones, see Fig.1) by comparing the 3'-tags. The unique cDNA species were assigned serial numbers from CELK00001 to CELK01508 .5'-tags of almost clones have been determined and database search showed that 582 species out of them gave significant similarities (blastx score > 100), which are listed below.(unavailable in WCS) This search has also detected many pairs of clones which appeared to be generated by alternative splicing. (b) Mapping: Thus far, 1,404 species have been mapped using the YAC polytene filters ( poly2 and sup poly2 );237 on LG1 ,244 on LG2 ,234 on LG3 ,245 on LG4 ,218 on LG5 ,210 on LGX and 26 on unassigned contigs. (c) Expression pattern: As a trial, 13 cDNA species were chosen at random and subjected to in situ hybridization on whole mount embryos (WBG 13-#2, p.22). The results were very striking that 6 species showed specific patterns of expression; yk3a 10(CELK00231) gave signals only in gut from the 2-fold stag, yk11 f3 (CELK00125) in several of ventral hypodermal cells at the comma stage, yk29 c5 (CELK01035) perhaps in P4 and its precursor cells, yk34 e2 (CELK01449) in several cells around nerve ring after the 2-fold stage, yk19 g11 (CELK00484) most cells except for perhaps P4 at the gastrulation period, yk35 c7 (CELK01166) in very early (<= 8 cell stage) embryos only. Being encouraged by the observation, we plan to start to survey expression pattern of individual clones using the in situ analysis. (2) Second set of cDNA clones: As the next step, we have picked up each 10,000 clones randomly from 3 different cDNA libraries; the same library of >2kb cDNA, a library of unfractionated cDNA, and an embryonic cDNA library. The total 30,000 clones were gridded and the cDNA clones belonging to the species which had been represented by more than 4 clones in the previous analysis were removed by hybridization screening with the 4+ cDNA probes. The resulting set of clones has been subjected to the above analyses although mostly 3'-tag sequencing so far, and we obtained 2,598 more clean 3'-tag sequences, which gave 1,204 more new cDNA species (see Fig.). The current status of our progress is that we have identified 2,712 unique cDNA species out of 5,792 clones (clean 3'-tags). 5'-tag sequencing and mapping on the genome are in progress.