Worm Breeder's Gazette 13(3): 53 (June 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Mutations in mua-3 cause a postembryonic failure of the attachments between the body wall muscles and the cuticular skeleton of the worm, resulting in a progressive paralysis of the affected animals. Bucher and Greenwald (Genetics 128:282-292) have shown by mosaic analysis that the focus of mua-3 activity is AB, presumably in the hypodermis, consistent with our observations of detachment in mua-3 animals in which the muscle bands remain intact, whereas the hypodermis ruptures. MUA3 may act as a link between the extracellular matricies (basal lamina or apically secreted cuticle) and the hypodermal cell or within the hypodermal cell in transmitting the contractile force across the hypodermal cell to the cuticle. In addition, the frequency of recovery of mua-3 alleles in genome wide screens for mua-class mutations suggests that mua-3 is a large gene.
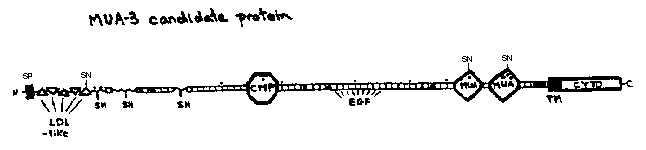
mua-3 maps between the previously cloned genes ced-7 and unc-69 ,within a 1.2 Mb region of chromosome III that has already been completely sequenced as part of the C. elegans genome project (WBG 12.2 pg14 -17).We inspected the predicted proteins identified by the sequencing project for potential candidates, based on our predictions for mua-3 size and function. The most promising candidate is that of hypothetical protein T20G5 .3.R. Durbin (personal communication) suggested that this protein might be only a portion of a larger one. Examination of the sequences upstream of T20G5 lead us suggest that T20G5 .3is probably only the C-terminus of a much larger protein, the N-terminus of which is identified as K08E5 .3by the sequencing project, and also containing some additional small EGF repeat containing ORFs between the two. In addition, we believe that K08E5 .3contains an erroneous frame selection by the Genefinder program, and should contain 5 rather than 3 LDL repeats. Our predicted protein of approximately 400 kd comprises 52 EGF repeats, 5 LDL receptor repeats, and a single cartilage matrix protein repeat, as well as a putative signal sequence and trans-membrane domain. Clearly it could serve as a linker between the hypodermal cell and extracellular matricies surrounding the hypodermis.
We obtained and injected only the cosmids containing this gene or its fragments, and obtained rescue with F55E6 a cosmid that physically contains the complete protein coding sequences. (The cosmid T20G5 that contains only T20G5 .3fails to rescue.) We are presently subcloning F55E6 to determine if the rescuing activity corresponds to the candidate gene or to a second gene contained on F55E6 .In addition, we are physically mapping the mua-3 mutations, to show that mua-3 is indeed this protein.