Worm Breeder's Gazette 13(3): 24 (June 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Proper cytoplasmic localization and asymmetric patterning are extremely important in animal development. The par genes are found to be involved in these processes during C.elegans early embryogenesis(1). In par-1 mutant embryos, first cleavage generates two nearly equal sized blastomeres; early cell divisions are synchronous; germ cell marker P granules fail to localize properly, and embryos arrest with an abnormally large number of fully differentiated cells. In addition, SKN-1 ,a putative transcription factor, thought to be a MS-cell fate determinant, is mislocalized in par-1 mutant embryos (2).
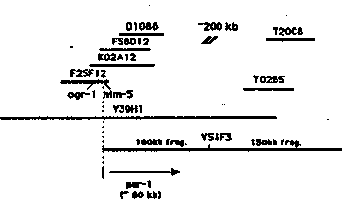
In order to understand the mechanism of cytoplasmic localization and cell fate determination, we seek to clone the par-1 gene. We mapped par-1 with respect to two cloned genes ogr-1 (3)and him-5 .Our data suggest that par-1 is located about 50-100 kb to the right of him-5 in a region of the physical map not covered with cosmids (Fig. 1). However, there are two YAC clones covering the gap (~200 kb). We performed germline transformation using DNAs made from these YACs, and one of the them, Y5 lF3 (~250 kb), rescued the par-1 mutant phenotype. We then subcloned Y5 lF3 into lambda. In the meantime, we found a restriction endonuclease AscI that cuts the YAC into 100 kb and 150 kb fragments (Fig. 1). Based on the mapping data, the 100 kb fragment is likely to contain par-1 ,so we used this fragment as a probe on a Northern blot which contains N2 and glp-4 ( bn2 )(germline deficient strain) mRNA, and identified a 4.4 kb maternally-enriched transcript. Since par mutations are strictly maternal, this germline enriched transcript is a candidate for par-1 . Subsequently, we identified a corresponding cDNA clone (named ZC22 )by screening Barstead's lambda-Zap cDNA library (thanks, Bob) with the 100 kb fragment. We then detected a par-1 allele associated polymorphism ( lw39 ,a gift from Jocelyn Shaw) with ZC22 .This result suggests that ZC22 is likely to be par-1 cDNA. In order to find a corresponding genomic clone to perform germline transformation, we used ZC22 as a probe to screen a C.elegans genomic DNA library (generously provided by Heidi Browning) and the phage libraries we previously constructed. Despite repeated attempts using various recombination-defective hosts, we failed to identify a genomic clone which could possibly contain the intact genomic region for the cDNA.
Because we are not able to perform germline transformation , rescue experiments with a par-1 genomic clone, we decided to determine if the cDNA we identified is par-1 gene by injecting into wild type worms antisense RNA made from this cDNA. People have injected anti-sense constructs and generated mutant phenotypes in C.elegans before (4), but to our knowledge direct injection of antisense RNA into syncytial gonad has never been tried. As a first attempt, we made in vitro transcribed anti-sense RNA from the cDNA clone and injected it into N2 worms. To our surprise, an average 50% of the embryos from the 16 injected worms arrested with par-1 terminal phenotypes (no morphogenesis; no intestinal cells; excess of pharynx). To convince ourselves that they were par-1 phenocopies, we analyzed the early cell divisions. The embryos have nearly equal first cleavage, synchronous early cell divisions and even showed a par-1 specific pseudocleavage defect. Besides par-1 characteristic embryonic lethality, we also observed that some viable progeny are sterile, which is consistent with the fact that a weak par-1 allele gives a grandchildless phenotype. As controls, we injected 1)4 N2 worms with DEPC- dH20 ;2)8 N2 worms with anti-sense RNA made from a Drosophila cDNA clone which encodes a cofilin protein; 3)8 N2 worms with anti-sense RNA made from a zygotic cDNA clone (identified by Jenny Watts in the process of cloning par-4 )into wild type worms, and we did not see any lethality. We are very interested in the anti-sense RNA injection result for two reasons: first, it indicates that the clone we identified is par-1 ;secondly, we think this technique is potentially applicable for other genes that are also in unclonable regions. We will try to find out if this technique will work for other maternal genes or possibly zygotically expressed genes.
We are in the process of sequencing the cDNA clone, and the sequencing information we have so far indicates that Par-1 is a putative serine, threonine kinase.
2) Bowerman et al., Cell Vol. 74, 443-452, 1993.
3) Abby Telfer thesis, unpublished.
4) Fire et al., Development 113, 503-515, 1991.