Worm Breeder's Gazette 13(3): 22 (June 1, 1994)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
The C. elegans transposons Tc4 and Tc5 were discovered because they are activated in mut-2 mutants [TR679 and its derivatives; genotype mut-2 ( r459 )]. The absence of evidence for movement in other backgrounds suggested that the transposition and excision of these elements might require the presence of mut-2 ( r459 ).We investigated this by analyzing reversion of unc-22 ::T c4 and unc-22 ::T c5 mutants in three different backgrounds: Bristol, mut-5 (aBergerac mutator), and mut-2 ( r459 ).In all cases, revertants were recovered only in the mut-2 mutant background, where they arose at high frequency (about 10+E-3). Molecular analysis of these revertants confirmed that nearly all resulted from excision of the inserted element from unc-22 .These results indicate that, as previously reported for Tc3 activity (WBG, Vol. 12 No.4, p. 18), Tc4 and Tc5 elements move only in the presence of the mutator activity encoded by mut-2 ( r459 ).
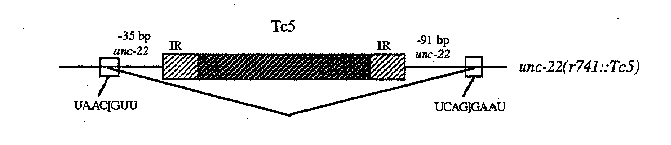
We are also investigating the fate of Tc5 sequences in unc-22 transcripts. Using RT-PCR, we are analyzing cDNA from several unc-22 ::T c5 mutants. Tc5 is spliced from unc-22 ( r741 ::T c5 )transcripts using donor and acceptor sites flanking the inserted element. The insertion site and splice sites are in the middle of a large unc-22 exon (these splice sites are not utilized in the processing of wild type unc-22 transcripts). The observed splicing removes all of Tc5 plus 126 bp of unc-22 coding sequence (see figure), generating a small, in-frame deletion in the mature mRNA. The twitching phenotype conferred by r741 may result from the loss of 42 amino acids from the encoded protein, or from inefficient splicing of Tc5 .Analysis of other unc-22 ::T c5 alleles is in progress.
We and others have shown that Tc1 , Tc3 , Tc4 and certain transposons in Drosophila, maize and humans are also spliced from transcripts of genes in which they are inserted. The demonstration that transposons can act as "mobile introns" has important implications. It suggests that the phenotypic consequence of a transposon insertion in a gene is determined not by insertion per se, but by how the transposon is spliced from pre-mRNA. It follows that many transposon insertions into coding regions may often be "silent". This idea is supported by our discovery that several revertants of unc-22 (r750:: Tc3 )still contain Tc3 in unc-22 (WBG, Vol. 12 No. 4, p. 16). As the C. elegans genome sequencing project proceeds, transposon-based methods to target mutations to specific genes will become more and more important. Those using such approaches should be aware of these features of C. elegans transposons.