Worm Breeder's Gazette 13(2): 14 (February 1, 1994)
These abstracts should not be cited in bibliographies. Material
contained herein should be treated as personal communication and
should be cited as such only with the consent of the author.
The C. elegans genome sequencing project: A progress report.
The C. elegans Genome Consortium, Genome Sequencing Center, Washington University School of Medicine, St. Louis, Missouri, USA and Sanger Centre, Hinxton Hall, Cambridge, UK.
We present here a progress report for the C. elegans genome sequencing
project. A detailed description of 2.181 Mb of finished, contiguous
sequence near the center of chromosome III has recently been accepted
(1). This sequence, bracketed by cosmid clones ZK112 and ZK757
,includes the mapped genes egl-45 ,
lin-36 , unc-36 , unc-86 , mig-10 , unc-116 , ceh-16 ,dpy- l9 , sup-5 , unc-32 , lin-9 ,gst-l lin-12 , glp-1 , emb-9 , tbg-1 and ncc-1 .All of the finished sequence data
for this region has been placed in acedb and the GenBank and EMBL
databases. Since the completion of the 2.181 Mb region, we have made
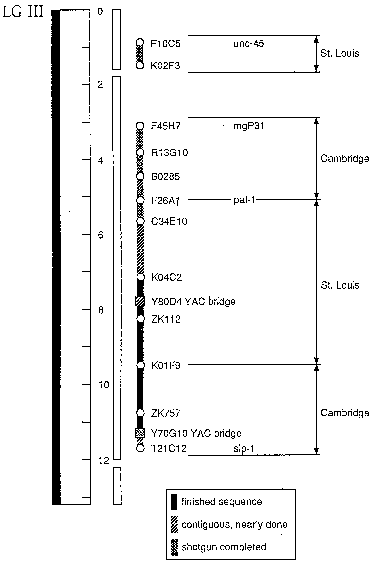
considerable progress on other large stretches of chromosome III (Figure
1). Several small regions not shown between F45H7 and T21C12 are
contained only on YAC clones and must be rescued and subcloned before the
entire sequence can be completed. Protein similarity data for a part of
the region is presented in Table 1 (database hits from the 2.181 Mb
sequence are not included in this list). The two groups have used
different criteria for determining which cosmids were included in the
list. Cambridge has included only finished cosmids and those cosmids
which are contiguous but still have one or more problem areas. St. Louis
has also included cosmids which have one or two gaps but which are
otherwise in good shape. Also, Cambridge has indicated the position and
type of similarity within each cosmid, while St. Louis has listed the
name and blastx score for the strongest hits. For future submissions we
hope to be more consistent; our experience here should help us decide
where to set boundaries for future Gazette releases. Although the cosmids
which contain database hits may not be complete, the Consortium will make
preliminary sequence data available to the community with the caveat that
it is preliminary and may still contain errors. Furthermore, we are
willing to help locate genes for persons having a bit of sequence data
(or to provide an estimated completion time for a particular cosmid). In
addition to chromosome III, sequencing has begun on chromosome II. Here,
we have started near the lin-5 gene,
with the St. Louis group proceeding left from cosmid C06A8 and the
Cambridge group proceeding right from cosmid T05A6 .For information on
homologies, please contact LaDeana Hillier (lhillier@watson.wustl.edu) or
Richard Durbin (rd@sanger.ac.uk). For information on sequencing plans or
estimated completion times, please contact Richard Wilson
(rwilson@watson.wustl.edu) or Alan Coulson (alan@sanger.ac.uk). All
requests for cosmid clones should be addressed to Alan Coulson.(1)R. Wilson et al. Nature, in press
Literature Cited:
1. R. Wilson et al. Nature, in press.
Figure 1