Worm Breeder's Gazette 13(1): 56 (October 1, 1993)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
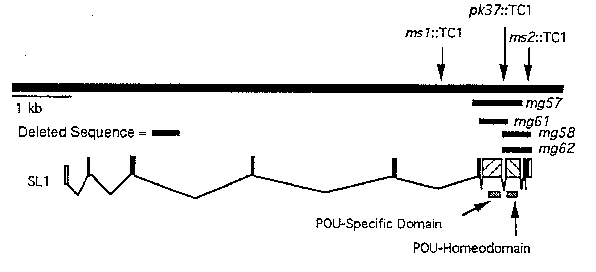
The POU-family of homeodomain-containing transcription factors has been implicated in controlling both cell type specification and cell proliferation during development. We have extended our previous studies of the POU-gene ceh-18 by using multiple TC1 -generateddeletions to analyze its in vivo function. We have obtained three TC1 insertions: msl:: TC1 is an insertion in a 2.6 kb intron just 5' to the POU-domain-coding sequences; pk37:: Tc1 is an insertion in the intron that lies between the coding sequences for the POU-specific region and the POU-homeodomain; and ms2:: Tc1 is an insertion in the last codon before the stop codon. The TC1 insertion strains do not have any obvious phenotypes upon observation with the dissecting microscope.
We have isolated 4 independent deletion alleles ( mg57 , mg58 , mg61 ,and mg62 ).We are still backcrossing the mutations, but we have a few preliminary observations. mg57 results from deletion of the entire POU-domain (POU-specific and POU-homeo domains) and has incompletely penetrant sterile and lethal phenotypes. Both of these were rescued by the cosmid C50A9 that contains ceh-18 .The L1 larval lethal phenotype of mg57 is similar to that which results from expression of antisense ceh-18 RNA from a heat shock promoter. We are interested in determining whether there is any relationship between the sterile mutant phenotype and expression of Ceh-18 protein by the distal tip cells in wild-type. mg58 and mg62 are unidirectional deletions that remove the homeodomain but leave the POU-specific region intact. mg58 and mg62 appear to have less severe phenotypes suggesting that the POU-specific region can mediate some level of function in the absence of a homeodomain. mg61 deletes the POU-specific region but leaves the homeodomain intact. This deletion is predicted to be out-of-frame if translation continues through the intron; however, because mg61 appears healthier than mg57 we are performing RT-PCR to determine if the extant splice donor is spliced to a cryptic acceptor.
We were surprised to find a few clustered mutations (CAAATTCTATT-->CAATTTCTATTT) in the intron sequence close to the 5' junction of the mg61 deletion. Perhaps this is due to error-prone repair. This was only observed in one of our four deletions.