Worm Breeder's Gazette 13(1): 30 (October 1, 1993)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
Transposable elements insert at many different positions in the genome, but the integrations are not necessarily randomly distributed over the genome. A detailed analysis of the insertion specificity of a transposon may be revealing with respect to the transposition mechanism of the element, since the integration preference may reflect how the integration complex interacts with the target sequence. We have investigated the target choice of the related transposable elements Tc1 and Tc3 .
The exact locations of 204 independent Tc1 insertions and 166 Tc3 insertions were determined. We used a PCR based approach which is sensitive enough to detect transposon insertions represented by a single DNA molecule and which does not involve a phenotypic selection for insertions (and is therefore unbiased for the distribution of insertions). The conditions were chosen such that insertions in an 1 kbp region of the genome were detected and that every insertion represents an independent somatic event.
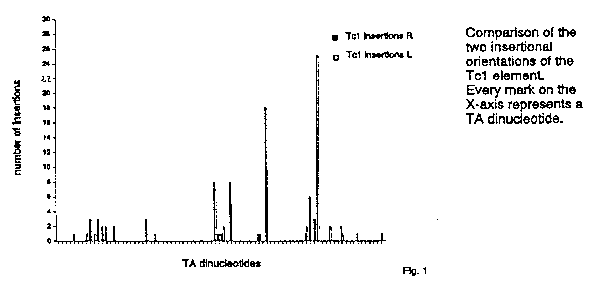
All 204 Tc1 insertions were into the sequence TA. Within the region investigated there are 82 TA's, but not every TA is used as often by Tc1 for integration (some TA's contain many insertions, others contain a few or no insertions; figure 1). The distribution of Tc1 insertions is independent of the orientation of the transposon. This implies that the integration reaction or the target sequence is symmetrical. We favour the first option since the flanking sequences of the TA's at the insertion sites have hardly any discernible symmetry.
We considered several explanations for the target site preference.
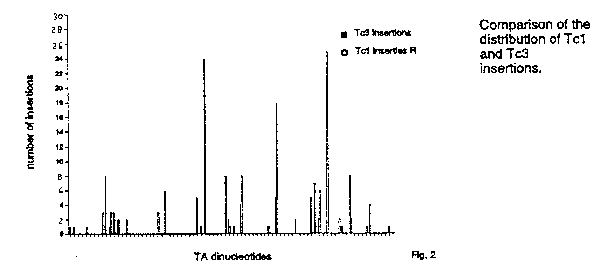
1) The local DNA structure. If only the local DNA structure makes one TA more attractive that another, we might expect to find a similar distribution for the two related elements Tc1 and Tc3 .Therefore we determined the exact locations of 166 Tc3 insertions. All insertions were into the sequence TA and they are also non-randomly distributed among the TA's in this region. We compared the distribution of Tc1 and Tc3 insertions and found that they are completely different (figure 2). Simple DNA structures (such as bends and kinks) are therefore not very likely determinants in target site selection (unless the two elements recognize them differently) .
2) The donor site. The non-random distribution of insertions could be explained if the original insertion site of the element effects the target choice. To investigate this we compared the distribution of Tc3 insertions to the distribution of a marked Tc3 transposon that jumped from one specific donor site in a transgenic line. The distribution of normal Tc3 insertions is the result of transposition from approximately 15 different donor sites. There is no difference between the two distributions (figure 3) indicating that the donor site has no effect on the target choice.
3) The flanking sequence of the TA dinucleotides. The most likely explanation of local target site preference is the recognition of the sequence flanking the TA dinucleotide by the incoming transposon. We investigated whether we could recognize a pattern of preferred sequences in the close vicinity of the insertion site. No clear consensus sequence was found. These studies focused on somatic transposition. Assuming these studies are also relevant for germ line transposition (and on the basis of germ line insertions studied thus far that seems a reasonable assumption) we do not think it is useful to look for "consensus" Tc1 sites when planning e.g. reverse genetic experiments.