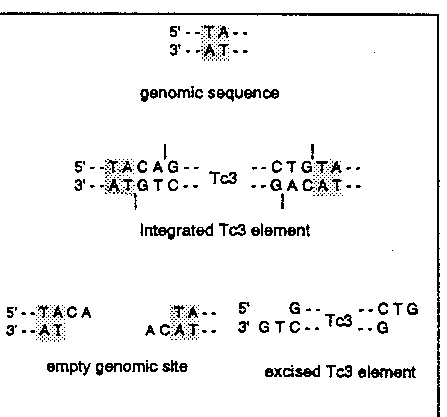Worm Breeder's Gazette 13(1): 29 (October 1, 1993)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
The transposable element Tc1 is thought to excises via double strand DNA breaks made at the ends of the inverted repeat (Plasterk (1991) EMB0 J. 7,1919-1925). Excised transposons have been detected as extrachromosomal structures (linear or circular; Rose and Snutch (1984) Nature 311, 485-486; Ruan and Emmons (1984) Proc. Natl. Acad. Sci. USA 81, 4018-4022). Excised, linear elements have also been detected for the related transposon Tc3 (Van Luenen et al. (1993) EMBO J. 12, 2513-2520). We have examined how the Tc1 and Tc3 transposons are excised by determination of the exact double strand cleavage sites.
Excision of Tc3 is not very frequent and thus the amount of extrachromosomal Tc3 in "natural" mutator strains is limited. Therefore we have made a stable transgenic line containing the Tc3 transposase under the control of an inducible promoter. Upon induction Tc3 transposase is made and this results in excision and transposition of Tc3 elements (Van Luenen et al. (1993) EMBO J. 12, 2513-2520). The level of excision and thus of the amount of extrachromosomal Tc3 elements is high.
The 3' end of the excised Tc3 transposon was determined in two independent ways. A restriction site in the inverted repeat located close to the end of the transposon was used to generate a small fragment containing the end of the transposon. This fragment and a size marker were run on a denaturing polyacrylamide gel. The DNA was Southern blotted and probed with a transposon end specific sequence. The length of the Tc3 fragment was compared to the size marker and from that we conclude that the 3' end of the excised element is the G nucleotide (see figure). Alternatively, the extrachromosomal material was amplified by PCR after 3' tailing with terminal transferase and subsequently the PCR product was sequence.
The 5' end of the excised Tc3 element was detected using the same Southern blot approach as used for the 3' end. The results indicated that the excised element ends at the G residue (see figure). In other words: the excised Tc3 element has a 2 nucleotide 3' extension. The same cleavage site is used most likely also for Tc1 excision. This has been observed in an in vitro cleavage reaction using recombinant Tc1 transposase. Cleavage was found directly after the G residue.
What are the implications of these results for the mechanism of transposition?
Since the sequences of Tc1 and Tc3 were determined it was unclear whether the TA dinucleotides flanking the integrated transposon are part of the transposon or result from target site duplication. Our results show that the TA dinucleotides are the result of target site duplication.
The element has to integrate using the 3' end, otherwise the last 2 nucleotides of the transposon will be lost. Integration through a nucleophilic attack of the 3' 0H group is also seen for other transposons. The presence of a 3' OH group at Tc3 ends is indirectly demonstrated by the possibility to add nucleotides to them by terminal transferase tailing.
Upon excision the cell is left with a double strand break with a 2 nucleotide 3' extension at both sides. The result of the cellular repair process is seen as a footprint left behind after excision of the element. We have sequenced somatic Tc3 footprints after PCR amplification. The footprints generated by Tc1 and Tc3 are very similar. The footprints are most often 4 basepairs in length (a TA basepair and two other basepair: CA or TG) and we conclude that these result from the 2 nucleotides at the 5' ends of the element which are not co-excised.