Worm Breeder's Gazette 12(4): 46 (October 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We are characterizing the pharyngeal muscle-specific enhancer from myo-2 with the goal of identifying factors controlling pharyngeal development. Our focus has been two enhancer fragments which can activate transcription in distinct patterns of pharyngeal cell types (1). A 150bp fragment we call the B element activates transcription in the major pharyngeal muscle cells m3 ,4, 5, and 7; while a 100bp fragment we call the C element activates transcription in all pharyngeal muscle cells and some non-muscle pharyngeal cells. We have previously suggested the B and C elements cooperatively activate transcription in response to different developmental signals.
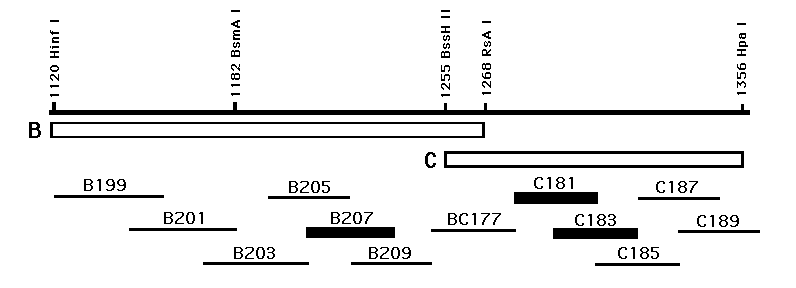
To identify cell type specific regulatory elements, we assayed double-stranded oligos spanning the B and C elements (numbered lines in figure) for enhancer activity. Three oligos (indicated by heavy bars) induce high levels of b-gal activity when cloned in multiple copies upstream of a myo-2 promoter :lacZ fusion. A 28bp oligo from the B element (B207) activates transcription in pharyngeal muscles nearly identically to the intact B element, and two overlapping oligos from the C element (C181 and C183 )activate transcription in pharyngeal muscle and nonmuscle similarly to intact C element. Mutations at the right and left ends of B207 and C183 eliminate activity while those near the center have no effect, suggesting at least two binding sites within the oligos are necessary for transcriptional activation. [See Figure]
We have used double-stranded B207 and C183 to screen a cDNA expression library for clones encoding site-specific DNA binding proteins. After screening approximately 400,000 recombinants, we have found 4 clones specifically binding B207 and 1 clone binding C183 .Of the clones binding B207 ,three are encoded by a new homeobox gene, designated ceh-22 .The ceh-22 homeodomain is most highly related (83% identical) to the NK-2 gene of Drosophila (2) and somewhat less highly related (60-73% identical) to additional homeodomains in Drosophila (2,3), humans (4), rats (5), and freshwater planarians (6). ceh-22 maps between her-1 and act-1 ,2, 3 on chromosome V in a region initially suggested by Thomas Burglin to contain a putative homeobox gene (7). The remaining clone binding B207 (provisionally named phage 21) encodes no obvious DNA binding motif but has features similar to both b-ZIP and b-hlh proteins. In preliminary experiments, we have failed to map phage 21 to a YAC.
DNA binding experiments with mutated oligos indicate the ceh-22 and phage 21 products bind to the left and right halves of B207 ,respectively. Therefore, both proteins may be necessary for myo-2 enhancer activity.
2. Kim, Y., and M. Nirenberg (1989) PNAS 86, 7716-7720.
3. Bodmer R. et al. (1990) Development 110, 661-669.
4. Price et al. (1992) Neuron 8, 241-255.
5. Guazzi, S. et al. (1990) EMBO J. 9, 3631-3639.
6. Garcia-Fernandez, J. et al. (1991) PNAS 88, 7338-7342.
7. Burglin T. et al. (1989) Nature 341, 239-243.