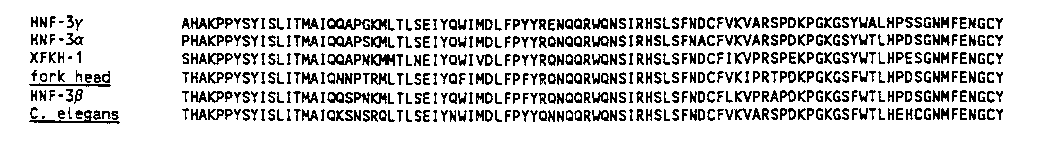Worm Breeder's Gazette 12(4): 32 (October 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
One approach to understanding gut development in C. elegans is to look for C. elegans homologs of genes involved in endoderm-specific gene expression in other organisms. We have focused our efforts on finding a C. elegans homolog of the Drosophila forkhead (fkh) and the rat HNF-3 (hepatocyte nuclear factor-3) genes since these genes appeared to be expressed in endoderm-derived tissues. HNF-3 (for which at least three different genes exist) is expressed in rat lung, intestine or liver, whereas fkh is found in different portions of the gut in Drosophila. These genes form a novel class of DNA-binding molecules and share a highly conserved 110 amino acid region (85% identity), which is believed to constitute part of their DNA-binding region. Recently, other genes sharing this highly conserved 110 amino acid domain have been identified in other organisms, such as the C. elegans lin-31 gene (Miller et al. WBG 12 (3), p.37). These related genes do not appear to be involved in endoderm development but rather to play roles in other tissues such as the brain (rat BF-1), the Spemann organizer (Xenopus XFKH1 ),or the vulva (C. elegans lin-31 ).It is therefore possible that only a subgroup of fkh/ HNF-3 -containingmolecules, such as those most closely related to the original fkh and HNF-3 ,are involved in gut development.
To clone a C. elegans fkh/ HNF-3 homolog, we designed a number of primers spanning the 110 amino acid region conserved between fkh and HNF-3 .These primers were used in the reverse transcriptase-PCR (RT-PCR), using total RNA obtained from a mixed population of N2 worms. The longest PCR product obtained (280 bp) was sequenced and found to encode a sequence of amino acids highly similar to the conserved fkh/ HNF-3 domain. Amino acid comparison between the Drosophila, rat, Xenopus and C. elegans molecules shows that they are 85% identical [See Figure 1]. This gene is also clearly distinct from lin-31 (59% identity).
We used this 280 bp PCR product to screen five genome equivalents of a C. elegans genomic library (partial S au3 Adigest in EMBL4 ;kindly provided by Chris Link) and 24 positives clones were isolated. All 24 clones appear identical on the basis of PCR, using primers spanning the conserved 110 amino acid region followed by digestion with diagnostic restriction enzymes. Sequencing of four of these clones confirmed that they are all identical and that .hey all contain the fkh/ HNF-3 domain. This has now been mapped by Alan and John to a rather empty region at the far right of chromosome V.
A 2.2 to 2.4 kb message expressed predominantly in embryos is detected upon analysis of a C. elegans developmental Northern using the 280 bp PCR product as a probe. We are currently sequencing the genomic clone and screening a cDNA library with the aim of making an antibody and looking for the expression pattern of this C. elegans fkh/ HNF-3 homolog during development. [See Figure]