Worm Breeder's Gazette 12(4): 27 (October 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
The glp-1 gene is required for at least two distinct cell-cell interactions during development: induction of anterior pharynx during embryogenesis and mitotic proliferation of germ cells during larval and adult stages. To identify other genes involved in these processes, recessive and dominant suppressors of glp-1 (ts) have been isolated. The 18 dominant suppressors, 8 of glp-1 ( q224 )and 10 of glp-1 ( q231 ),are tightly linked to glp-1 and may be intragenic revertants or mutations in a nearby gene, e.g. lin-12 .These suppressors are not simply revertants of the original glp-1 (ts) mutations because some Glp worms are seen in each suppressed strain.
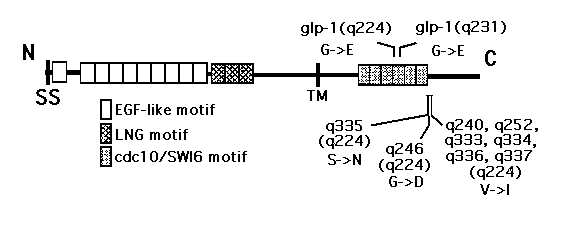
We are using SSCP to determine whether dominant suppressor mutations are located within glp-1 . Genomic DNA from each suppressor strain is arnplified by PCR using glp-1 specific primers in the presence of a-32P-dATP. Eight overlapping primer pairs yielding 700-1400 bp fragments span the glp-1 gene. The labelled products are digested with restriction enzymes, denatured, and fractionated on non-denaturing 6% acrylamide/10% glycerol gels. The mobility of single-stranded DNA molecules is determined by conformation, which is sequence dependent, and length. Fragments containing a single base substitution can adopt a conformation different from wild type and can migrate with altered mobility in a non-denaturing gel. We have performed SSCP analysis of the entire glp-1 gene from all 18 dominant suppressors. Seven of the q224 suppressors had unambiguous fragment shifts; sequencing of unlabelled PCR products revealed a single base change within each shifted fragment [See Figure]. No shifts were detected for q246 ;however, given the close proximity of the other q224 suppressors, we sequenced the corresponding region in q246 and did find a base change. All of the changes found so far are G-C -> A-T transitions (as expected for EMS-induced mutations), cause amino acid changes, and are clustered near the original q224 mutation. Interestingly, six of the q224 suppressors are identical even though they were found in four separate screens. We plan to recheck the original freezer stocks to confirm this finding. The q224 mutation is located in the fourth cdc10 /SW16repeat in the putative intracellular dornain of glp-1 and the suppressors flank the end of the sixth repeat. cdc10 /SW16repeats have been shown in other systems to mediate protein-protein interactions. Thus, we propose that the q224 change disrupts a crucial protein-protein contact and that the dominant suppressor mutations partially restore the structure and allow appropriate protein-protein interactions. The bad news is that we have not yet found any of the q231 suppressors. We plan to test different SSCP conditions on these alleles and to partially sequence some as well.