Worm Breeder's Gazette 12(4): 16 (October 1, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
In the course of analyzing footprints generated by germ-line excision of Tc3 from unc-22 we have isolated four independent revertant alleles that still have intact (almost, see below) Tc3 elements at the insertion site. As described in an accompanying article (Collins, Mills, Reynolds and Lyons; WBG: this issue) we isolated and analyzed a collection of independent, spontaneous revertants of TW241 [genotype unc-22 ( r750 ::T c3 ); mut-2 ( r459 )].To determine for each revertant allele the nucleotide sequence of the expected "empty site" generated by the assumed excision event, we amplified genomic DNA via PCR using unc-22 primers flanking the insertion site, then sequenced the PCR product. Most revertant alleles analyzed had lost Tc3 ,with a small footprint remaining at the excision site. However, four revertant alleles, cj206 , cj213 , cj221 ,and cj222 ,were not that simple. Our first clue: repeated efforts to amplify the empty site region failed. Our next clue came from a closer look at the phenotype of these revertants. Each has a "partial" revertant phenotype: animals twitch slightly upon exposure to nicotine but do not twitch detectably in its absence. [ unc-22 mutants twitch violently in the presence of nicotine; wild-type animals are paralyzed in the same conditions. Most, but not all, unc-22 mutants twitch in the absence of nicotine as well.]
We tested a straightforward explanation for these observations: that the revertant alleles have lost flanking unc-22 sequences in addition Tc3 .For the C. elegans transposon Tc1 ,4 of 51 revertants of one unc-22 ::T c1 allele exhibited partial revertant phenotypes, and three of these had deletions of 1-2 kb of flanking sequences (Kiff et al. Nature 331:631-633). Deletions of this nature would explain our inability to amplify the empty site region. Total genomic Southern blots shot down this idea. To our surprise, 3 of 4 revertants had blot patterns indicating Tc3 was still present at its original insertion site. The fourth allele appeared to have retained the Tc3 element plus an additional 400 bp. Of course, one of us (guess who) assumed this reflected a mix up in strains by another one of us. Careful repetition gave the same result. Before yielding to the evidence we tested an obvious prediction: if Tc3 is still present we should be able to amplify the insertional junctions. We tried and we did. For each allele, amplification with appropriate primers (one in Tc3 ,one in flanking unc-22 sequences) yielded products for both junctions, and the sizes were consistent with the Southern blot data. I gave up: Tc3 is still present in these four alleles.
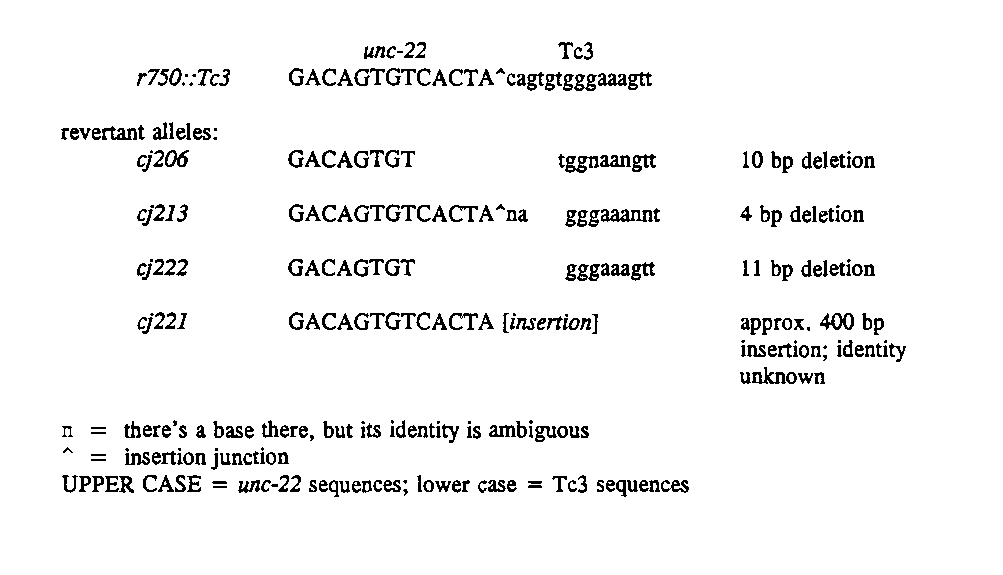
We were faced with four cases of reversion of an unc-22 ::T c3 allele without excision of Tc3 from its site of insertion. How can that be? Extragenic suppressors are not responsible for the revertant phenotype: backcrossing each revertant with N2 failed to resegregate F2 twitchers. In search of a molecular basis for these events, we looked at the nucleotide sequences of the amplified insertional junctions. This work is still in progress, but the emerging picture is that each revertant allele contains a disruption at the upstream junction; the downstream junction is unchanged (polarity is with respect to the unc-22 transcript). The figure below shows the nucleotide sequences of the upstream junction of unc-22 ( r750 ::T c3 )and the four partial revertants.[See Figure]
Two questions arise:
(1) What's going on? How do these changes restore unc-22 gene function with an almost intact Tc3 element still in the gene? Our working hypothesis is that they in some way alter the splicing of the unc-22 transcript such that Tc3 sequences are spliced out, or spliced out differently compared to r750:: Tc3 .We are looking at the appropriate region of the unc-22 transcript in the mutant and revertants to determine if the r750:: Tc3 transcript contains Tc3 sequences, and if the revertant transcripts are different. Understanding the basis of the revertant phenotype for these alleles will provide insight into the larger question: do all transposon insertions in genes cause mutations. If not, what factors determine whether or not a particular insertion will have phenotypic consequences. [For more in this question see the article by Rushforth and Anderson in this issue of WBG].
(2) How common is reversion without excision? Four of 17 revertants of unc-22 ( r750 ::T c3 )arose without loss of Tc3 .We have not detected any such events for Tc5 [0125 for unc-22 ::T c5 and mec-7 ::T c5 (from Savage and Chalfie) revertant alleles] and are not aware of any for Tc1 among nearly a hundred revertants of Tc1 -inducedalleles of many genes that have been analyzed. Perhaps it is unique to Tc3 .Perhaps it is strongly dependent on flanking sequence context or proximity to splice sites, or other less apparent factors. The work of Engels and co-workers on P elements in Drosophila demonstrated that transposon excision does not always result in reversion of the mutant phenotype caused by the insertion. In fact, their results indicate that the vast majority of excision events have no phenotypic consequences because the process that repairs the double-strand gap at the excision site uses an insertion-containing template on the sister chromatid or homologous chromosome (Engels et al. Cell 62:515-525). The inserted element is "copied back in" . In this light, the results of Mori et al. (Mol. Gen. Genet. 220:251-255) suggest the same is true for Tc1 ,as subsequently demonstrated by Plasterk (EMBO J. 10:1919-1925) and by Moermann et al. (NAR 19:5669-5672). So transposon excision without reversion may be common. Our observations indicate that reversion without excision is also possible. Further work will reveal how common it might be for Tc3 ,and Tc1 ,2,4,5,6.
Mori et al., Mol. Gen. Genet. 220:251-255.
Plasterk, EMBO J. 10:1919-1925
Moermann et al., NAR 19:5669-5672.