Worm Breeder's Gazette 12(3): 74 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
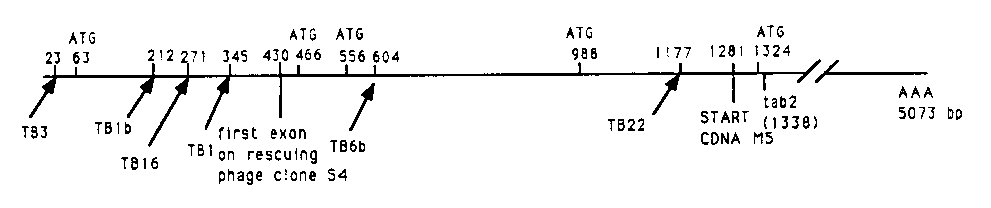
In the last worm meeting I reported the further phenotypic characterization and the cloning of the unc-53 gene. To find the 5' end of the unc-53 transcript I did nested PCR on L2 stage random primed cDNA (gift from D. Zarkower), between oligo tab2 [See Figure 1] from the 5' end of the original unc-53 cDNA M5 and an oligo to the SL1 transspliced leader sequence. This yielded at least 6 classes of PCR-fragments which have been subcloned and sequenced. All contain the 43 bp between oligo tab2 and the 5' end of cDNA M5 ( bp1281 to 1338 in fig.). Much of the cDNA between basepairs 430 and 1338 maps to the 16.4 kb lambda clone (54) that rescues the Unc and E g1 phenotypes of unc-53 ( n152 ).This and additional PCR reactions with various internal primers suggest that the PCR reactions extended the correct transcripts.
The longest PCR fragment (TB3; see figure) extends the sequence of cDNA M5 with 1280 bp. When added to the length of the cDNA M5 ,this unc-53 transcript would then be 5073 bp long (including some polyA tail) and have a 1528 AA open reading frame. I probed a Northern blot (also a gift from D. Zarkower) of L2 , L4 and adult poly(A)+ RNA with cDNA M5 .This identified a major 5.0 kb transcript (based on position relative to the large tra-1 transcript) and at least 2 smaller transcripts. This (preliminary) experiment shows that unc-53 is expressed in L2 , L4 and adult worms, and suggests that little (if any) of the unc-53 5'- end is missing. The smaller PCR-fragments TB1 b, TB16 , TB1 , TB6 band TB22 are "nested deletions" of clone TB3 with SL1 'sat their 5' end. The sequence of each is identical in the regions of overlap. The shorter SL1 transspliced transcripts contain ATG's downstream of the SL1 addition sites at positions 466, 988 and 1324. Comparison to the sequence of genomic clones confirmed that the positions where SL1 is added in TB22 and TB6 Bare intron-exon boundaries. Position 430 identifies an intron-exon boundary onto which no SL1 was found spliced indicating that not all splice acceptor sites receive SL1 .
The alternative unc-53 5'-ends may arise by the insertion of SL1 on multiple intron-exon boundaries of a single unc-53 transcript. Alternatively the unc-53 gene may have different promotors for each isoform. A similar phenomenon was described for sdc-1 by Nonet and Meyer (Nature Vol. 351, page 65-68,1991). They find only one trans-spliced form and showed using primer extension that more than 90% of the sdc-1 messages are not transspliced. It needs to be
examined whether the unc-53 5' ends reported here are made in vivo and encode different proteins or whether they represent PCR noise.
Unc-53 is a large gene with at least 18 kb between the first and last exon of the 5 kb message. The shortest lambda clone that rescues does not contain the first 430 bp of the unc-53 transcript. This suggests that the ORF between pos. 63 and 430 is not essential for transformation rescue. This rescue could thus come from expression of transcripts TB6 bor TB22 or from "non-specific" initiation of transcription on the extrachromosomal arrays.
Searches of the databases with the complete unc-53 ORF have not yielded credible homologies or clues about its cell biological function. Using an internal unc-53 methionine as a start codon, I over-expressed and purified a 47 kd part of unc-53 in E. coli. One affinity purified rabbit serum, 4 rat polyclonal sera and one rat monoclonal antibody (16-48-2) work at high (1:30.000) titres on Western blots of the 47 kd unc-53 fragment expressed in E.coli. These five reagents fail to detect antigen of the correct size on Western blots of total worm proteins or worm proteins fractioned by progressive extraction with detergents, urea and SDS. They also fail to give a consistent, credible staining pattern in immunofluorescence of embryos and adults. It is therefore unlikely that unc-53 is expressed at high levels. Over-expression of unc-53 cDNA in C. elegans or mammalian tissue culture cells may be required to identify where in the cell unc-53 acts.