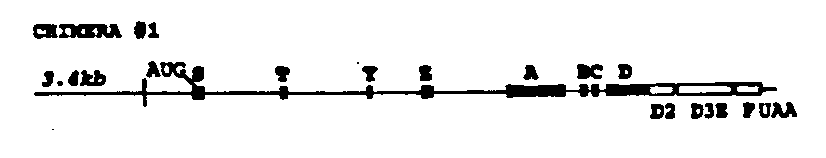Worm Breeder's Gazette 12(3): 66 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
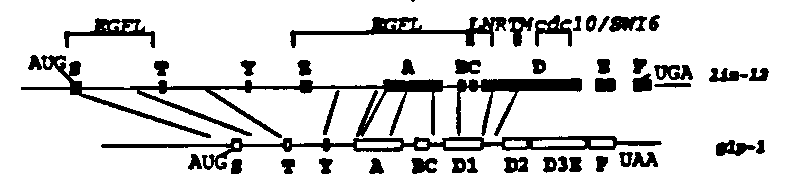
We are currently trying to establish the relationship of various lin -12 structural motifs to their function in vitro. As a first step in this process we have attempted to map functional domains of the lin -12 protein using lin 12/ glp-1 protein chimeras under lin-12 regulation.
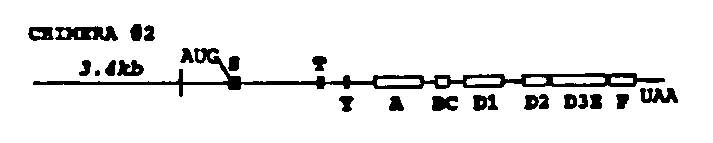
Two chimeras were constructed (see the figure below) by fusing corresponding introns or exons of lin-12 and glp-1 (Yochem and Greenwald Cell 58: 565, 1989; Austin and Kimble Cell 58: 565, 1989). Each construct contains 3.4 kb of lin-12 5' flanking sequence, which is sufficient to allow a lin-12 genomic DNA clone to complement a lin-12 null mutation (Wilkinson and Greenwald WBG 12: 61, 1992). Each construct was injected at a concentration of 6ug/ml along with a dominant rol-6 marker (Mello et al, EMBO J 10: 3959, 1991) at lOOug/ml into N2 wild type hermaphrodites. Arrays from the resulting Roller lines were then crossed into a lin-12 (n676 n909a m)background and assayed for rescue of the Lin-12 (0)sterility and protruding vulva defects.
In each case the chimera containing array was able to complement lin 12( n676 n909 ).These results indicate that lin-12 / glp-1 chimeras most likely will not be useful for mapping functional domains of lin-12 (or glp-1 ).However, these results are interesting in that they strongly support the hypothesis of Lambie and Kimble (Development 112: 231, 1991), who suggested that lin-12 and glp-1 are functionally interchangeable based on the novel "Lag" phenotype of a lin-12 (0) glp-1 (0) double mutant. (Additional support for this hypothesis was also provided by Mango et al, Nature 352: 811, 1991.)
We have now begun a structure/function analysis of the lin-12 protein by making specific deletions and point mutations, starting with deletions of various EGF like repeats in the extracellular portion of the protein.
Austin and Kimble Cell 58: 565, 1989
Wilkinson and Greenwald WBG 12: 61, 1992
Mello et al, EMBO J 10: 3959, 1991
Lambie and Kimble. Development 112: 231, 1991
Mango et al, Nature 352: 811, 1991