Worm Breeder's Gazette 12(3): 54 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
How many ras proteins are there in C. elegans? This has been a common question asked to us since the let-60 gene was shown to encode a ras protein. A high degree of identity in overall structure and a substantial functional overlap may be required for a ras-like small GTPase to be called ras. In worm, loss-of-function in let-60 ras abolished its roles in vulval cell fate specification as well as in early larval growth. It is thus unlikely that C. elegans has another ras protein which has the redundant functions of let-60 ras. Similar words can be said about the Drosophila ras-1 gene. A few years ago, three Drosophila "ras " genes were identified, but ras-2 and ras-3 differ in overall structure from ras-1 and mammalian H-ras significantly and may only be called ras-like small GTPase. The Drosophila ras-3 has been renamed rap-1 .
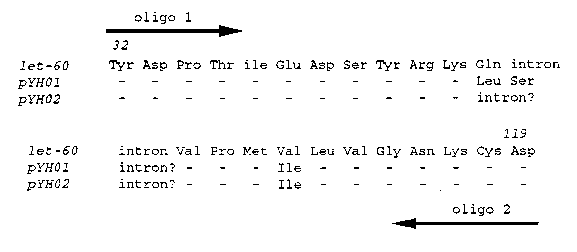
Some years ago, ras homologs in C. elegans were searched for with a mammalian ras gene as probe. Two candidates were identified and named ras-1 and ras-1 (marked somewhere in the physical map), even though they are not very homologous to ras. To investigate the roles of other ras-like small GTPase during C. elegans development, we started to search for let-60 ras homologs. Since ras-like proteins are conserved in their GTP/GDP- binding domain and the so called "effector domain", we used degenerate oligonucleotides homologous to these regions to PCR probe the genomic DNA. Although all ras coding region are similar in size, their intron sizes are expected to be different. Our initial screen detected three bands which differ in size from let-60 .The DNA fragments within the two primers were cloned. The partial sequence of two of the clones are shown below. Both genes are highly homologous to let-60 ras in the two small regions. The genomic DNAs containing the fragments have been isolated from a phage library from Bill Wood's laboratory. We have not been able to isolate cDNAs for the two genes after screening nearly a million plaques out of the Barstead cDNA library. The RNA of these two genes (if they are real genes), could be far less abundant than that of let-60 .We will continue to characterize these two candidate genes and hope to study the functions of ras-like small GTPases by molecular genetic methods including constructing dominant mutations in vitro.