Worm Breeder's Gazette 12(3): 47 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
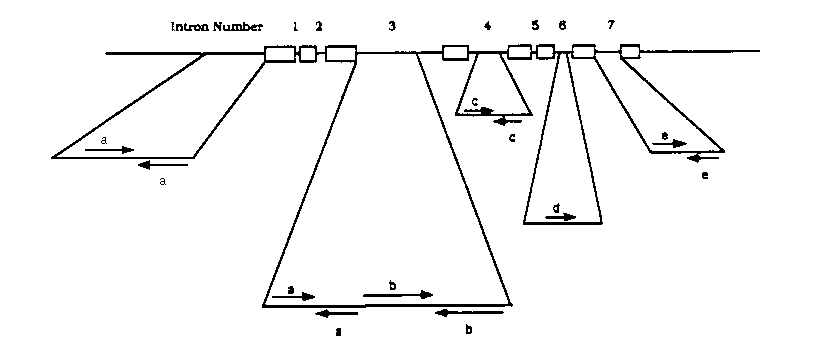
The ced-3 gene is necessary for the proper execution of programmed cell death in C. elegans. Recessive mutations in ced-3 block most programmed cell deaths. The gene has been cloned and cDNAs have been sequenced (J. Yuan, personal communication). The putative Ced-3 protein encodes a novel protein of 503 amino acids with a highly serine-rich amino-terminus. To characterize the ced-3 locus further we determined the sequence of a 7.5 kb genomic fragment capable of rescuing ced-3 mutant animals and found that the ced-3 gene includes eight exons and seven introns. All putative splice sites fit the C. elegans consensus splice sequences. Four of the introns, as well as the region 5' to the putative start codon, contain several repetitive elements (see article this issue by Shaham et al. and figure below). Element a in intron 3' and in the 5' upstream region resembles sequences found in the fem-1 and hlh-1 genomic sequences. Elements c and d in introns 4 and 5, respectively, are found in the genomic lin-12 sequence and in two sequenced cosmids (B0303 and ZK643 ).Two other elements (b and e) are novel.
To confirm that the message encoded by the 7.5 kb genomic fragment is the ced-3 message, we sequenced exons and exon/intron boundaries from 12 ced-3 mutants. We found alterations in all 12 mutants. Eight mutants contained missense mutations, three contained nonsense mutations, and one has a T instead of a conserved G in a splice acceptor. Nine of the 12 mutations are clustered in the three carboxy-terminal exons, two are in the first exon and one in the fifth. There are no mutations in the serine-rich region, suggesting either this region is unimportant or individual residues are functionally redundant.
To see if any of the mutations affect the level or size of the ced-3 message, we analyzed RNA from the 12 ced-3 mutants by Northern hybridization. All 12 mutants contained a 2.9 kb RNA corresponding to the wild-type message in length. In addition, no significant alteration in the ced-3 message steady-state level was apparent. That the putative splicing mutants show no alteration in message length or level is surprising, but might be explained by the presence of cryptic splice sites.
Finally, we used a combination of primer extension and amplification of ced-3 RNA by PCR to define the 5' end of the ced-3 message. Our results suggest that at least some of the ced-3 message is spliced to SL1 at a splice site 65 bp upstream of the putative start codon.