Worm Breeder's Gazette 12(3): 38 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
We are continuing our analysis of the post-transcriptional regulation responsible for the dramatic reduction of lin-14 protein levels during the L1 stage. High levels of lin-14 protein specify early cell fates while low levels of protein specify later cell fates. Two lin-14 gain-of-function mutants fail to down-regulate lin-14 protein levels at post-L1 stages, resulting in the reiteration of L1 and L2 cell fates at later stages. Both gain-of-function mutations are disruptions of the 31JTR of the lin-14 gene, suggesting that sequences in the 3'UTR are normally responsible for mediating the down-regulation of lin-14 protein. Because lin-4 (1p) mutants are similar to lin-14 (gp) mutants at the level of phenotype and effect on lin-14 protein regulation, the lin-4 gene may play a direct role in down-regulating lin-14 protein levels (see Wightman et al, Genes Dev., 1991).
We are currently testing the sufficiency of the lin-14 3'UTR to mediate temporal down-regulation. Transcriptional fusions of the col-10 promoter to the lacZ coding region with either the unc-54 , lin-14 ,or lin-14 ( n536 )deletion 3'UTR have been constructed and introduced into worms along with the rol-6 marker. In stable array strains bearing the unc-54 3'UTR construct, X-gal staining is observed in hypodermal cells at high levels in late embryos and all post-embryonic stages. Animals bearing constructs with the lin-14 3'UTR in place of the unc-54 3UTR displayed detectable X-gal staining only in late embryos, L1 and early L2 animals. Animals bearing constructs with a lin-14 3'UTR which is deleted for the same 607 bp deleted in the n536 mutant display bright X-gal staining at all stages. Finally, when the construct which utilizes the intact lin-14 3'UTR is crossed into a lin-4 ( e912 )mutant background, bright X-gal staining is now observed at all stages. These results indicate that the lin-14 3'UTR may be sufficient to confer temporal regulation on col-1 O-drivenlacZ expression, and that lin-4 exerts its effect on lin-14 protein levels via the lin-14 3'UTR. This result is particularly interesting in light of the possibility that lin-4 encodes an active RNA molecule (see Feinbaum, Lee, Ambros, last WBG issue). The lin-4 RNA could interact with the 3'UTR of the lin-14 mRNA, leading to a change in the processing, export, or translation of the lin-14 message at later stages. The fusion constructs are being integrated into the chromosome to control for copy number and allow a quantitative description of the effect.
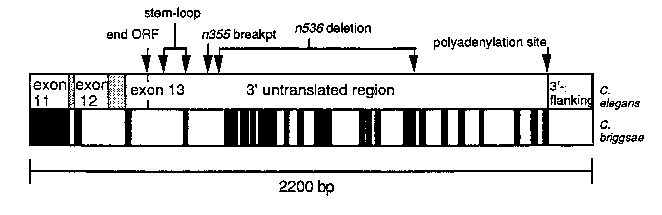
In order to delineate which regions of the 1.6 kb lin-14 3'UTR may be required for this regulation, we have cloned and sequenced the lin-14 3'UTR from C. briggsae to identify sequences that are conserved between the two species. Stretches of exact sequence conservation up to 32 bp long are found throughout most of the 3'UTR. Significantly, there is little primary sequence conservation between the species from the 3' end of the ORF to the 5' breakpoint of the n536 deletion, while within the region deleted in n536 there are numerous extensive regions of sequence conservation. These sequences may represent cisacting sites to which other factors, perhaps lin-4 ,bind and mediate down-regulation, or they may represent regions that are conserved so as to maintain a secondary structure required for normal lin-14 regulation. We have also found that the position of a predicted favorable stem-loop structure is also conserved between C. elegans and C. briggsae, although the primary sequence of the stems and loop structure are not conserved. Black boxes on the lower line of the figure below show the locations of conserved sequences between the two species over a 2.2kb portion of the 3' end of the gene. The upper line of the figure shows landmarks in the C. elegans sequence. Splice donor and acceptor sites are conserved, yet the predicted C-termini of lin-14 proteins are not conserved at the nucleotide or amino acid level. These data suggest that the C-terminal 65 amino acids of the C. elegans lin-14 protein may not be essential for function.