Worm Breeder's Gazette 12(3): 35 (June 15, 1992)
These abstracts should not be cited in bibliographies. Material contained herein should be treated as personal communication and should be cited as such only with the consent of the author.
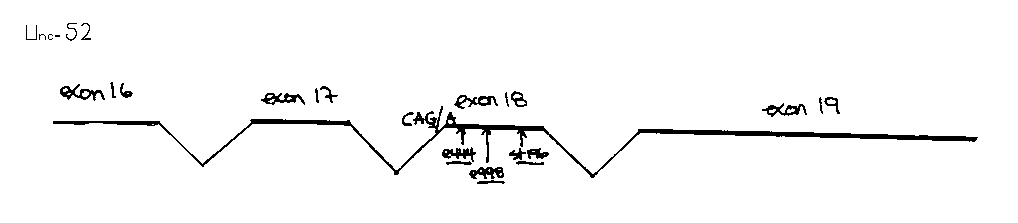
The unc-52 gene spans almost 15 kilobases of genomic DNA and produces at least three alternatively spliced transcripts (WBG Vol.12, No.2, p.50). Several different classes of unc-52 mutation have been identified, based on the mutant phenotypes they produce and their complementation patterns. Class 1 mutants develop normally as larvae, but adult animals are usually paralyzed except for their heads. Alleles of this category include e444 , e669a m, e998 , e1021 , e1421 and st196:: Tc1 .Class 2 and class 3 alleles result in larval lethality. The class 2 allele, st549 ,fails to complement all other unc-52 alleles. In contrast, the class 3 allele, ut111 ,complements the class 1 alleles.
We have identified the molecular lesions responsible for four unc-52 alleles. We examined DNA from most of the mutants and about 30 intragenic revertants using PCR analysis. All of the unc-52 mutants gave wildtype sized PCR fragments except for two. The st196 allele contained a Tc1 element inserted into exon 18 as previously determined (T. Rogalski and D. Moerman, unpublished results). The class 3 allele, ut111 ,had also been isolated from a mutator strain (by K. Kondo in I. Katsura's lab), but had a very different phenotype from that exhibited by the st196 allele. PCR analysis using Ben Williams Tc1 -mappingprimer (which recognizes the inverted repeats of this element) along with primers from within unc-52 indicated that ut111 contained a Tc1 insertion into exon 2. Curiously, only unc-52 primers 3' to the insertion gave amplification products in combination with the Tc1 primer. The inverted repeat at the 5' end of the element may, therefore, be altered, although the element does appear to excise somatically. We're currently trying to sequence the 5' end of the element to determine why the primer doesn't work for PCR amplification in this direction.
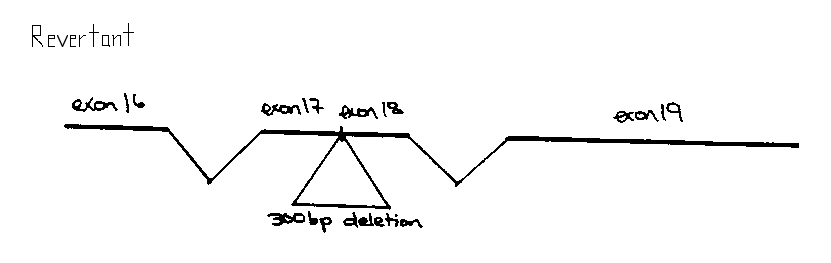
Although 29/30 of the intragenic revertants gave apparently wildtype sized fragments upon PCR analysis, one e998 revertant proved to have a 300bp in frame deletion, thus fusing exons 17 and 18 (see figure). Sequencing of this interval indicated that the e998 mutation was a G-A transition that resulted in a premature TGA stop codon in exon 18, just upstream of the st196:: Tc1 mutation. Subsequently, we have determined that the e444 mutation is also a premature TGA stop codon approximately 1OObp upstream of e998 .Not all class 1 mutations are in exon 18. Although we have not yet found the lesions responsible for the e669 , e1021 ,and e1421 mutations, this region is wildtype in these animals.
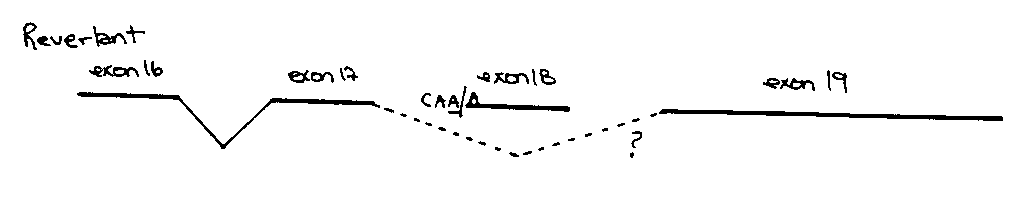
We have also sequenced the e998 revertants which gave wildtype sized fragments upon PCR analysis. Two spontaneous revertants (provided by Teresa Rogalski) have exact reversions of the e998 mutation back to wildtype. Five other revertants were isolated in EMS screens and all show a change from CAG/A to CAA/A in the splice acceptor site at the beginning of exon 18. The G which has been changed in these revertants is normally found at the 3' end of all C.elegans introns and is thought to be essential for splicing (WBG Vol . 12, No . 2, p . 57 ) . Presumably, elimination of this exon containing the e998 mutation, results in a wildtype phenotype. Splicing of exon 17 directly to exon 19 would not change the reading frame of the message [See Figures]. We are currently trying to do PCR analysis of cDNA from these revertants to determine which alternate splice acceptor sites are being used.
A technical note: We have been using Ben William' s protocol for PCR analysis on single worms (we use two worms). We then use 1/25th of this product, without purifying at all, for sequencing using Molly Craxton's linear amplification technique (the BRL kit).
WBG Vol . 12, No . 2, p . 57